Taxa Name
| Taxa
Level | Barcode
Data? | Number of
Barcodes
from
any ocean | Number of
Barcodes
from this region | Barcode Locations and
Species Occurence Map | MZGdb Atlas
v2025-m10-01
2025-Oct-27 |
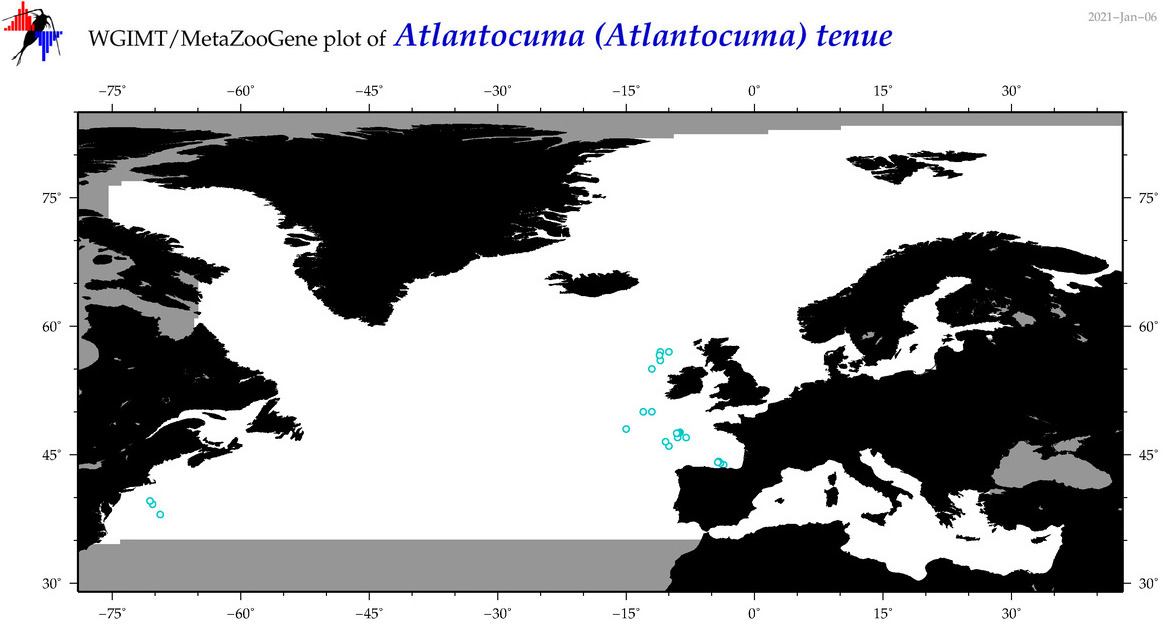
| Atlantocuma (Atlantocuma) tenue |
Species
(1) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094802
R:1:0:0:0 |
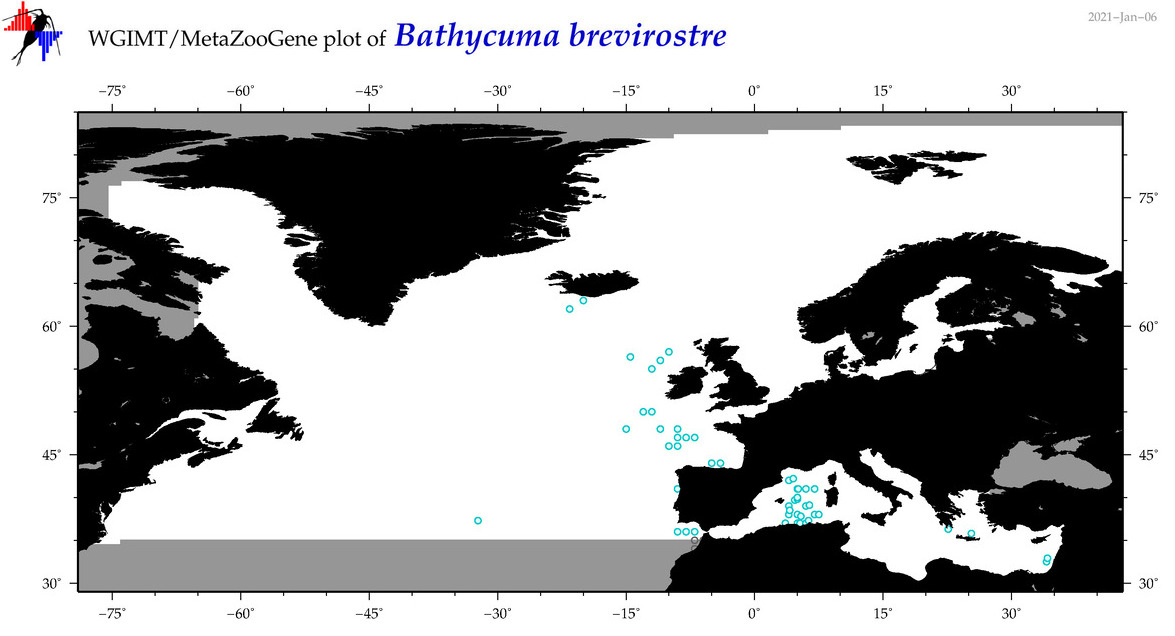
| Bathycuma brevirostre |
Species
(2) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4045194
R:1:0:0:0 |
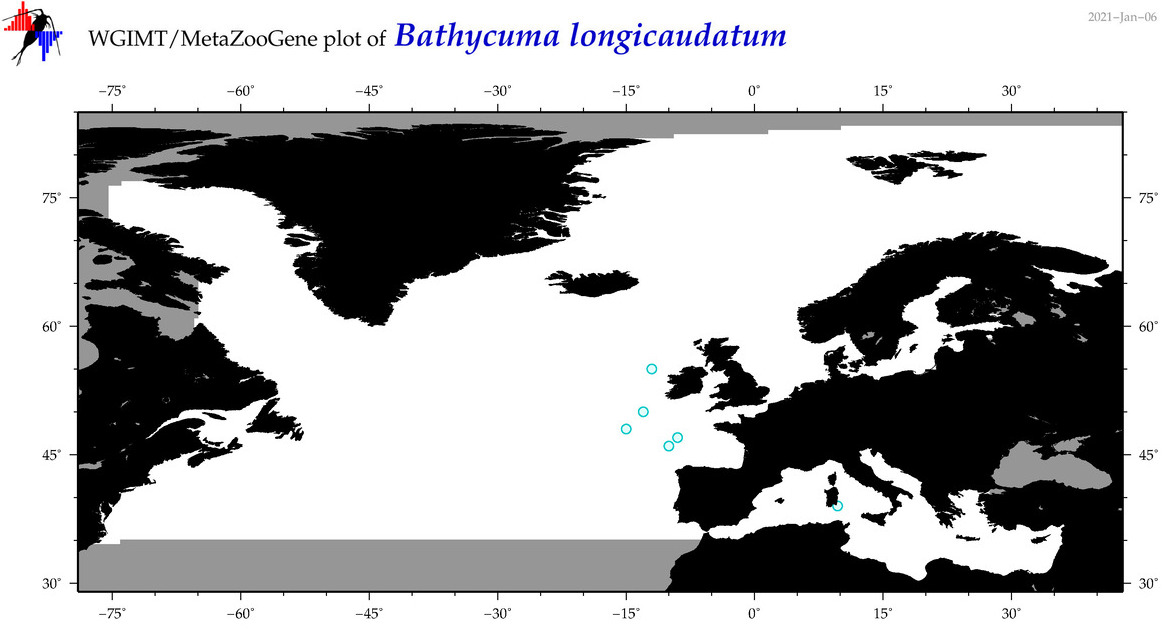
| Bathycuma longicaudatum |
Species
(3) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4045197
R:1:0:0:0 |
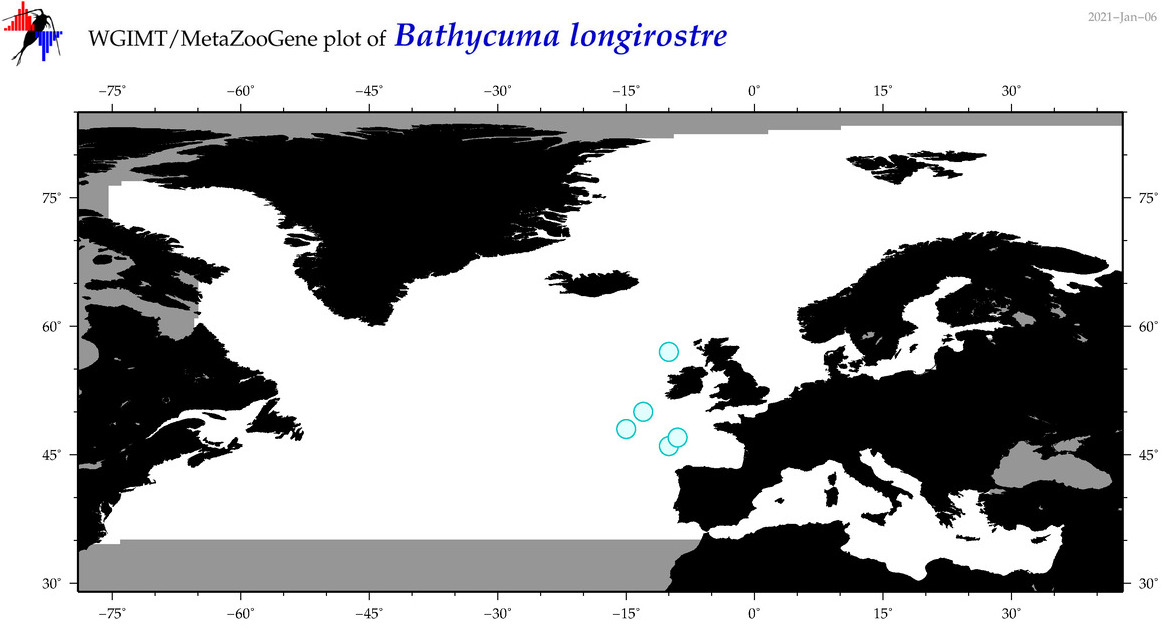
| Bathycuma longirostre |
Species
(4) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4045198
R:1:0:0:0 |
| Bodotria arenosa |
Species
(5) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
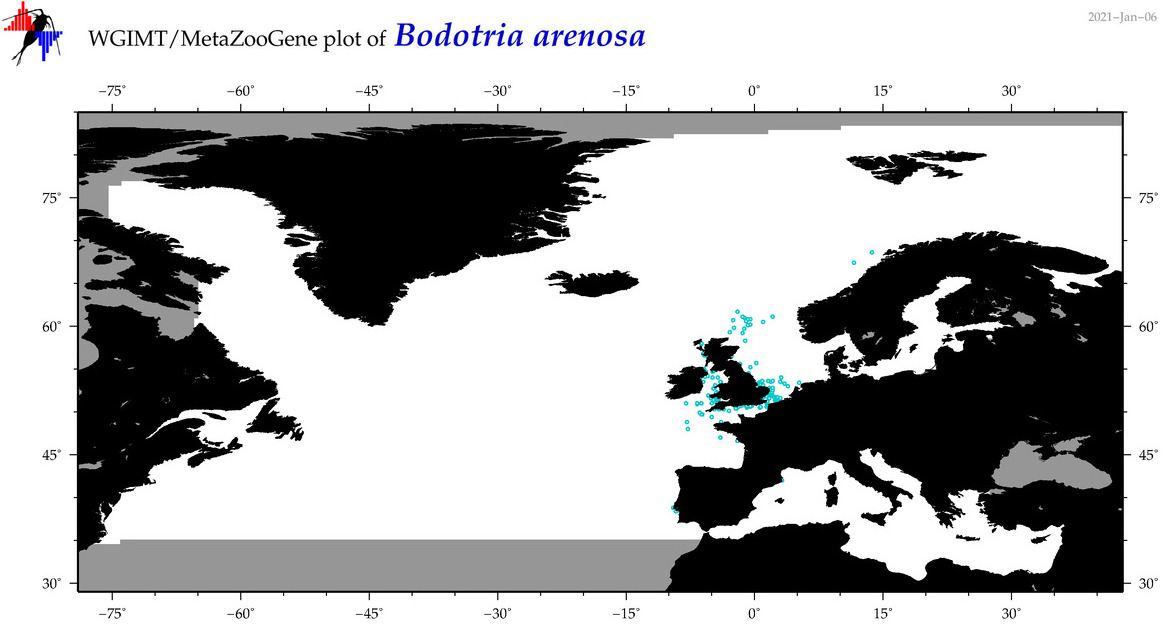 |
accepted
T4045619
R:1:0:0:1 |
| Bodotria pulchella |
Species
(6) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
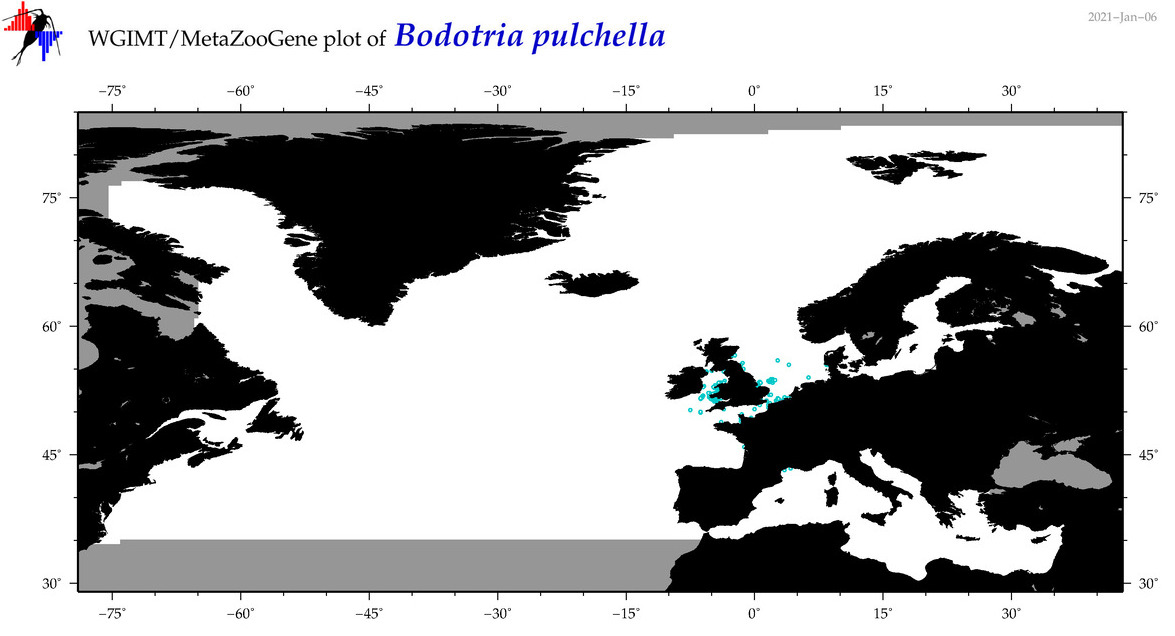 |
accepted
T4045633
R:1:0:0:0 |
| Bodotria scorpioides |
Species
(7) |
COI
12S
16S
18S
28S
|
COI = 11
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
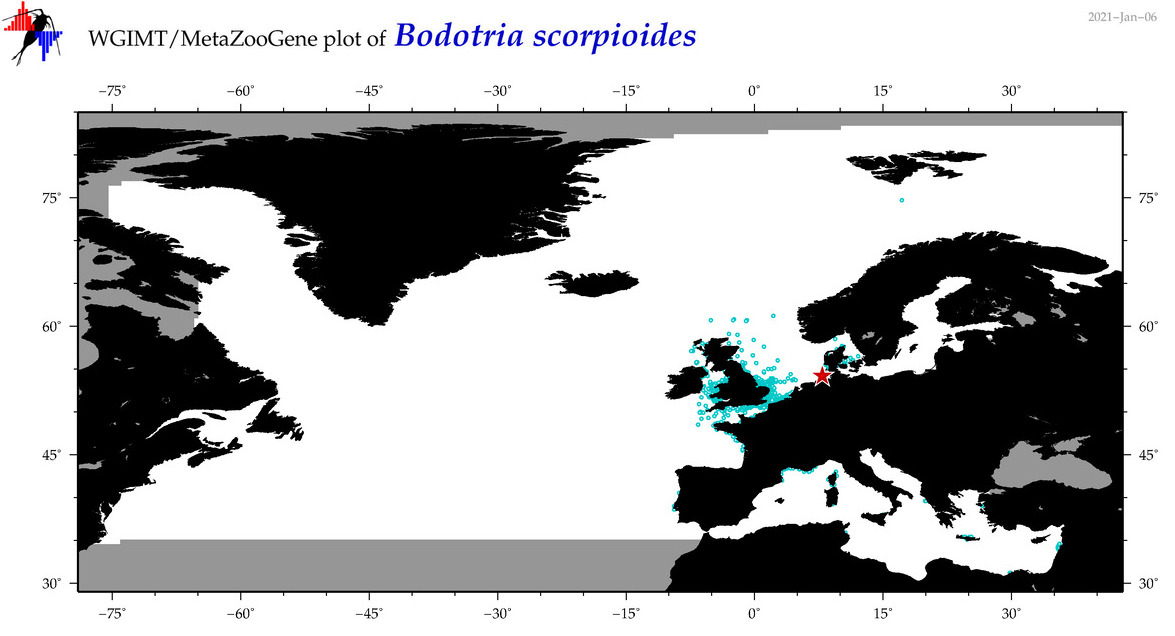 |
accepted
T4010328
R:1:0:0:0 |
| Brachydiastylis resima |
Species
(8) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
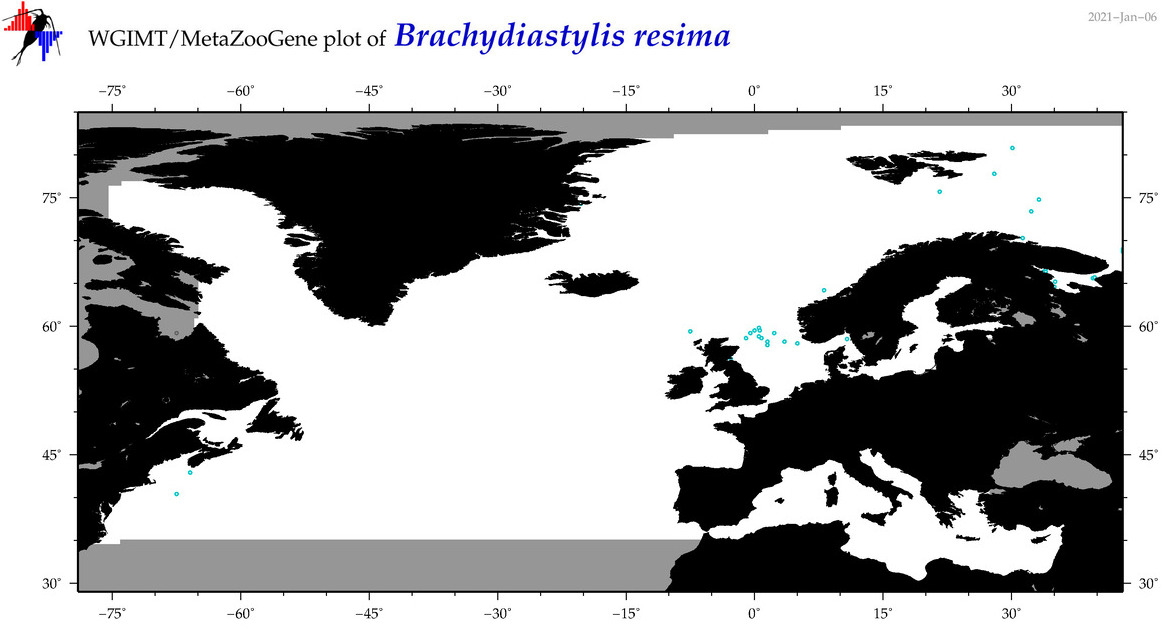 |
accepted
T4045779
R:1:1:0:0 |
| Bytholeucon hiscens |
Species
(9) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046102
R:1:1:0:0 |
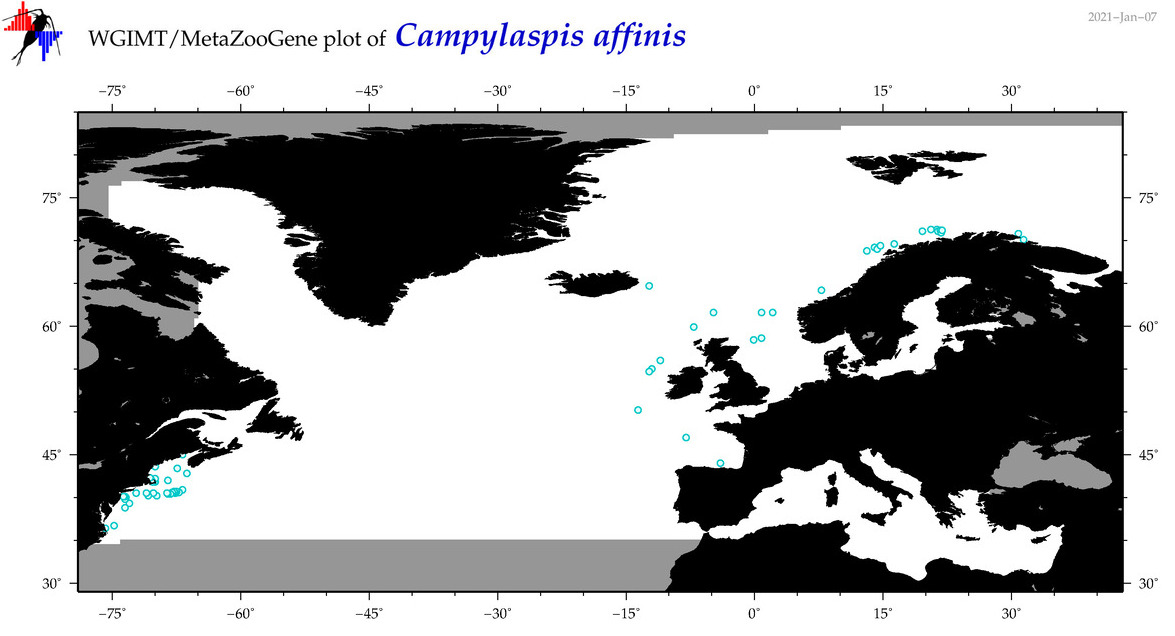
| Campylaspis affinis |
Species
(10) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046523
R:1:0:0:0 |
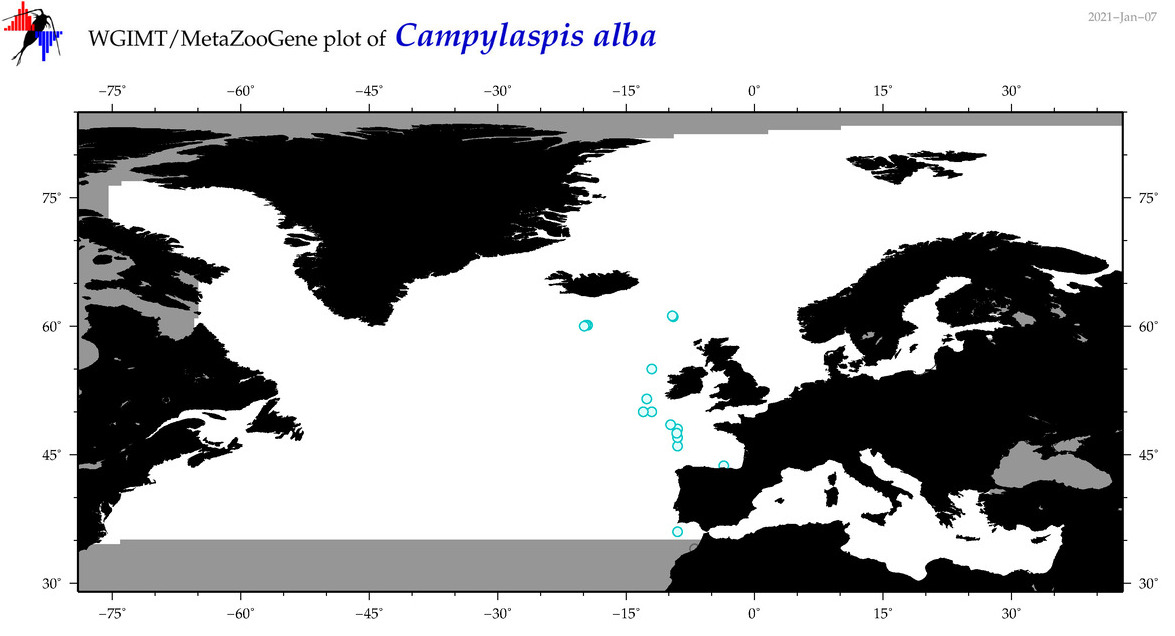
| Campylaspis alba |
Species
(11) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046525
R:1:0:0:0 |
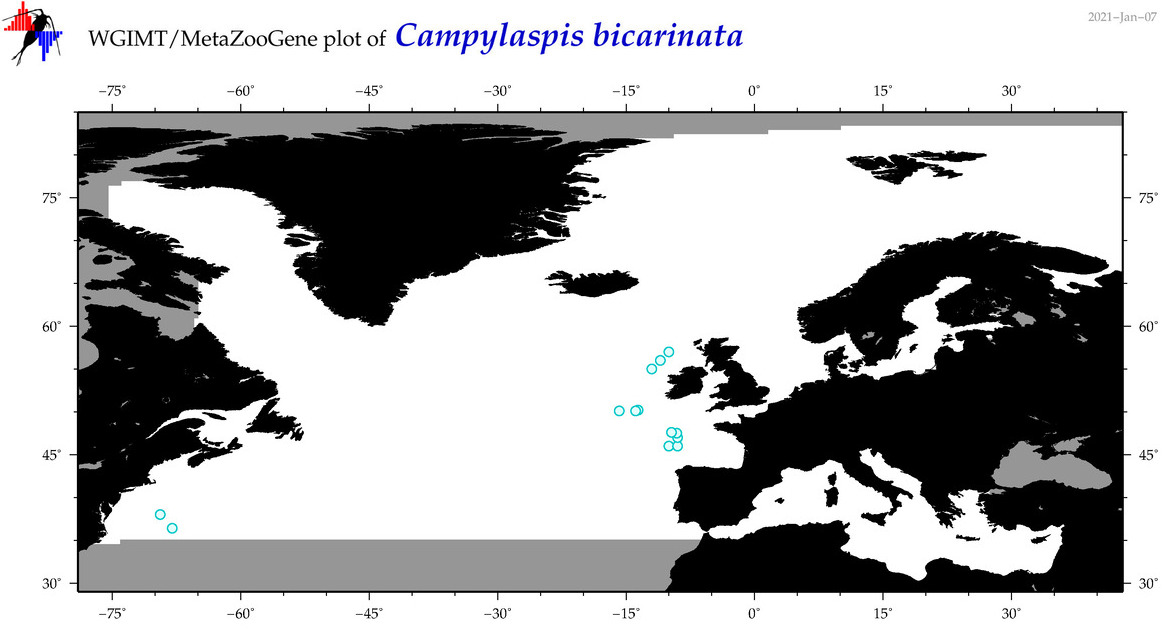
| Campylaspis bicarinata |
Species
(12) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046533
R:1:0:0:0 |
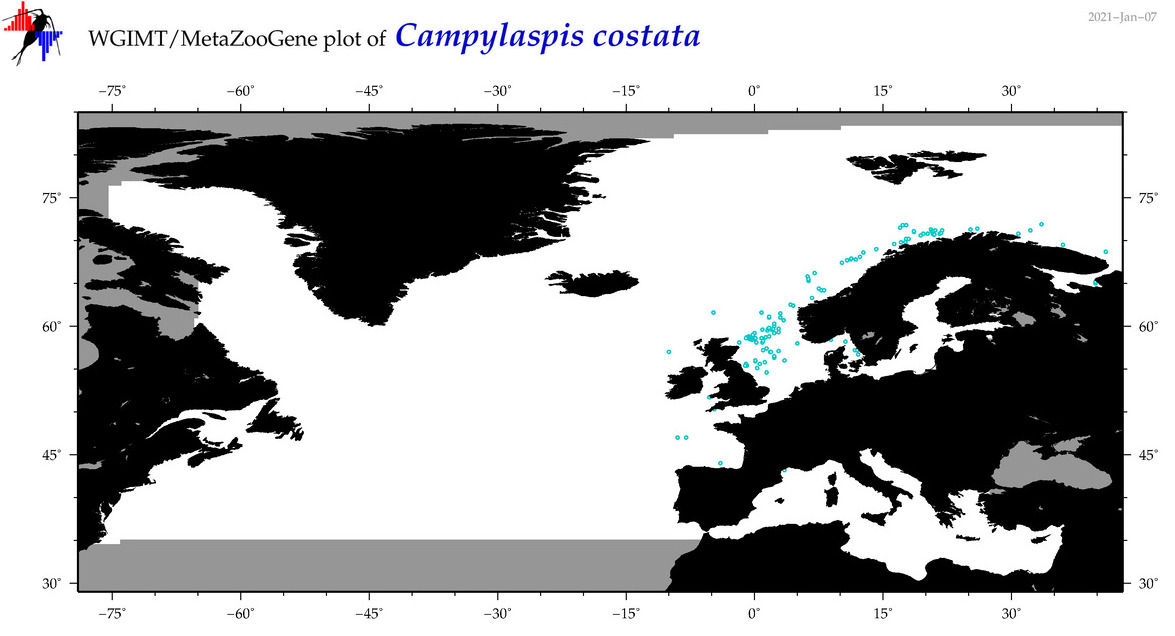
| Campylaspis costata |
Species
(13) |
COI
12S
16S
18S
28S
|
COI = 5
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046544
R:1:0:0:0 |
| Campylaspis glabra |
Species
(14) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
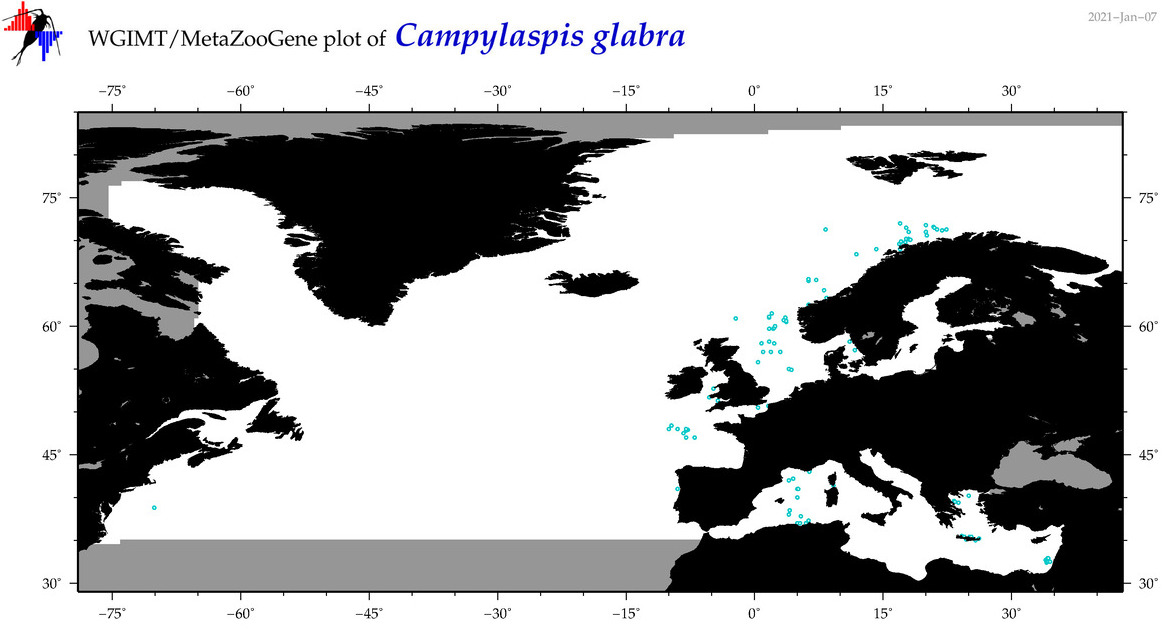 |
accepted
T4046554
R:1:0:0:0 |
| Campylaspis globosa |
Species
(15) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046556
R:1:0:0:0 |
| Campylaspis horrida |
Species
(16) |
COI
12S
16S
18S
28S
|
COI = 5
12S = 0
16S = 1
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
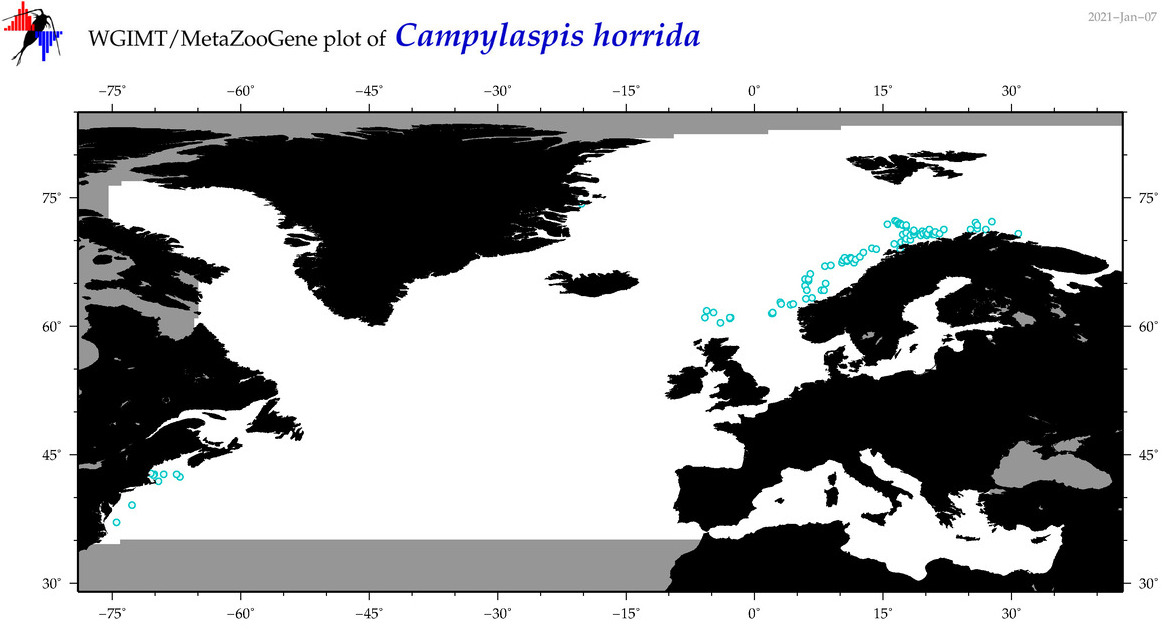 |
accepted
T4046565
R:1:0:0:0 |
| Campylaspis legendrei |
Species
(17) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
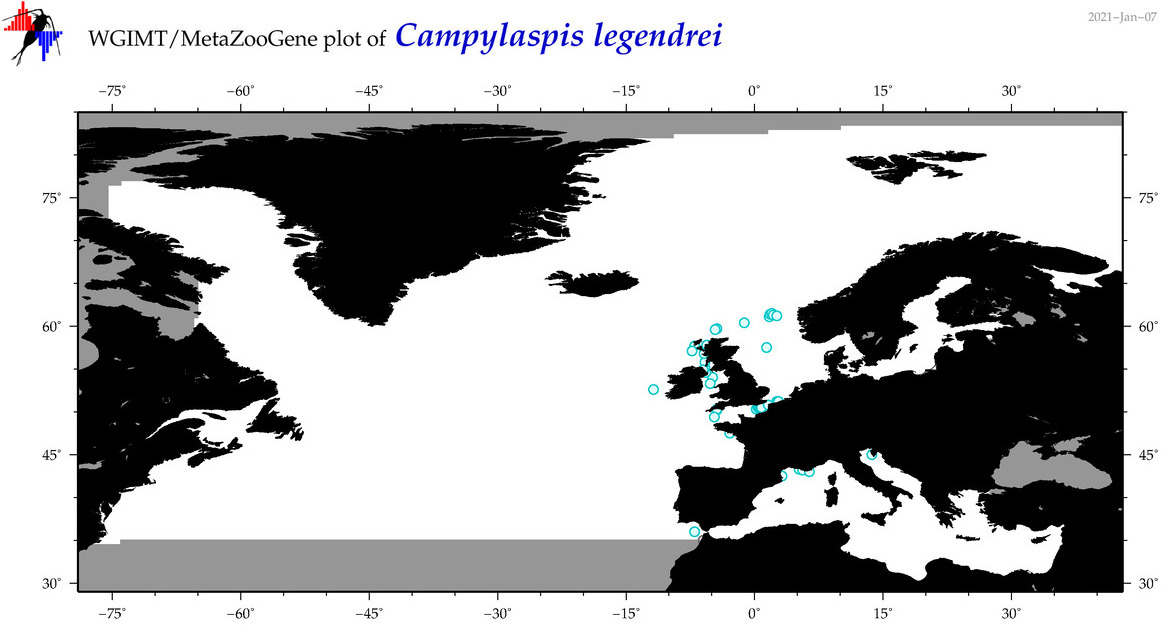 |
accepted
T4046577
R:1:0:0:0 |
| Campylaspis mansa |
Species
(18) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046584
R:1:0:0:0 |
| Campylaspis nitens |
Species
(19) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
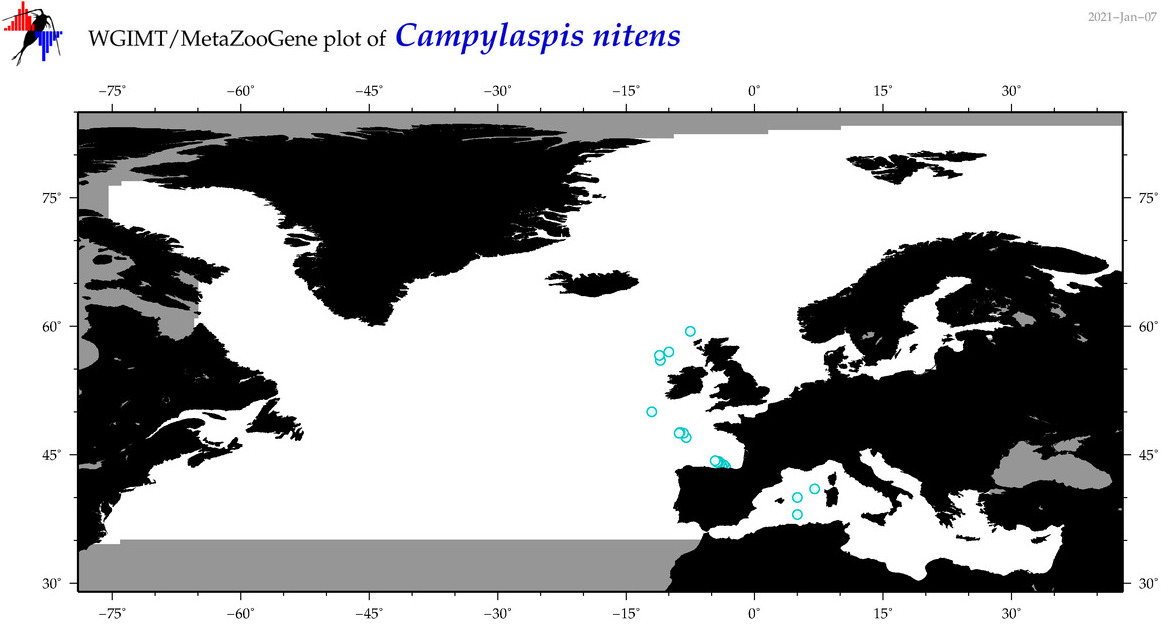 |
accepted
T4046592
R:1:0:0:0 |
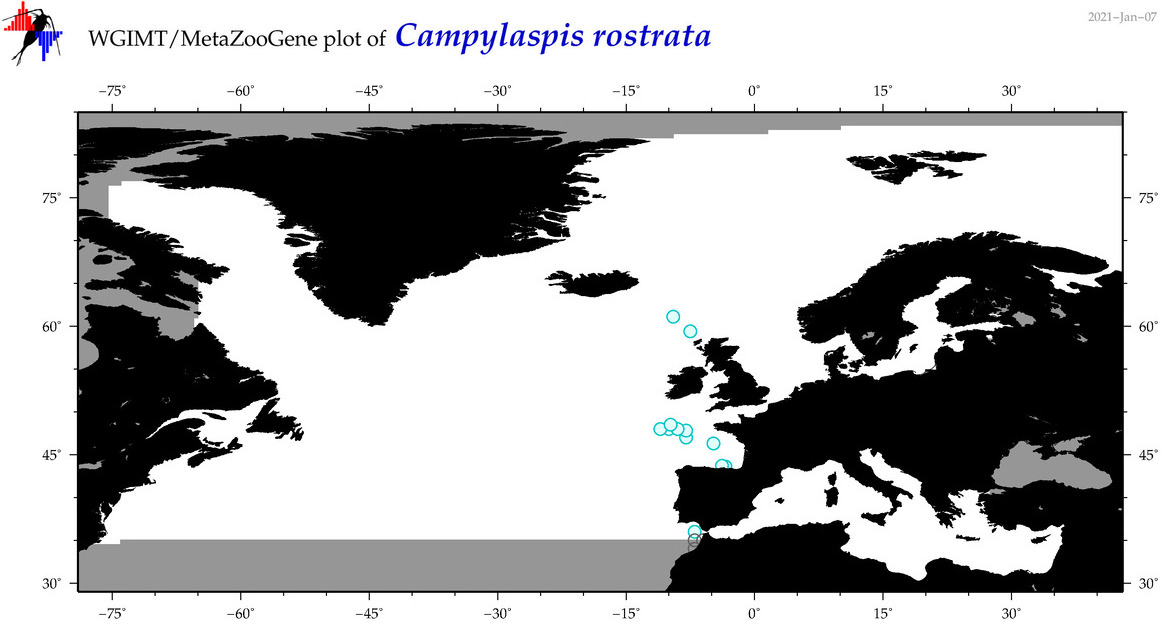
| Campylaspis rostrata |
Species
(20) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046614
R:1:0:0:0 |
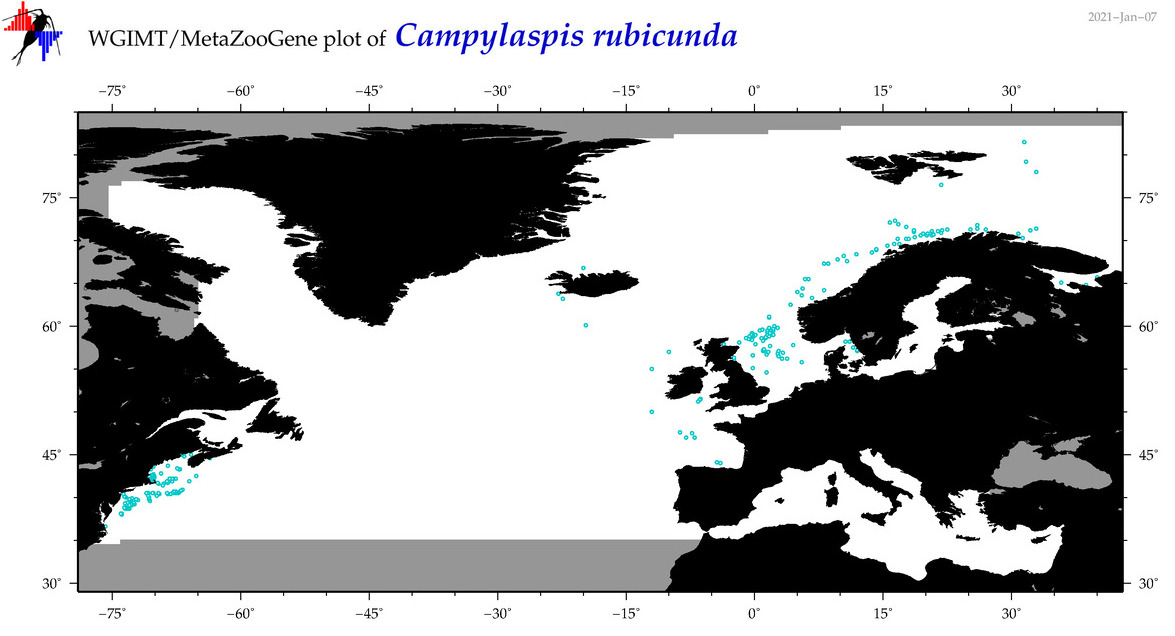
| Campylaspis rubicunda |
Species
(21) |
COI
12S
16S
18S
28S
|
COI = 2
12S = 0
16S = 7
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046615
R:1:0:0:0 |
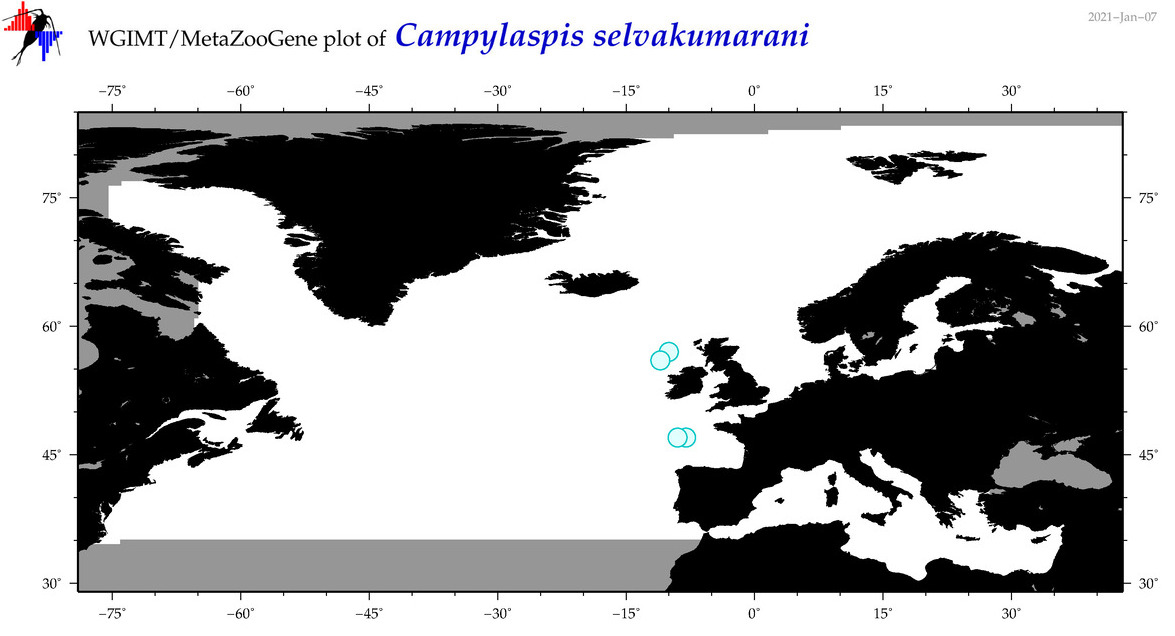
| Campylaspis selvakumarani |
Species
(22) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046622
R:1:0:0:0 |
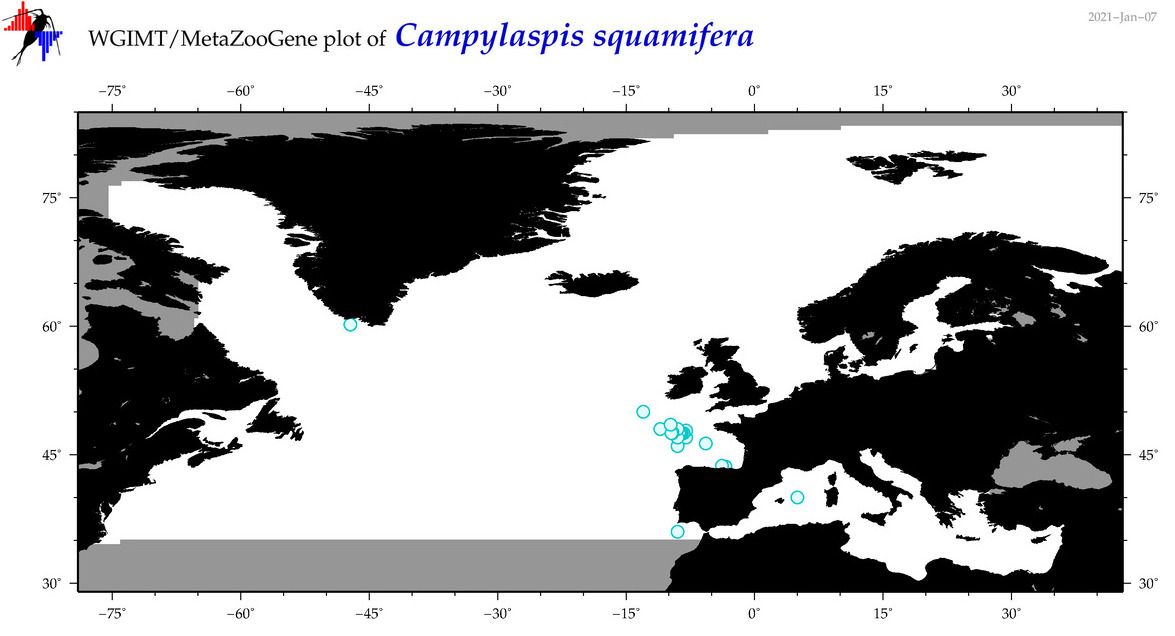
| Campylaspis squamifera |
Species
(23) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046629
R:1:0:0:0 |
| Campylaspis sulcata |
Species
(24) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 3
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046631
R:1:0:0:0 |
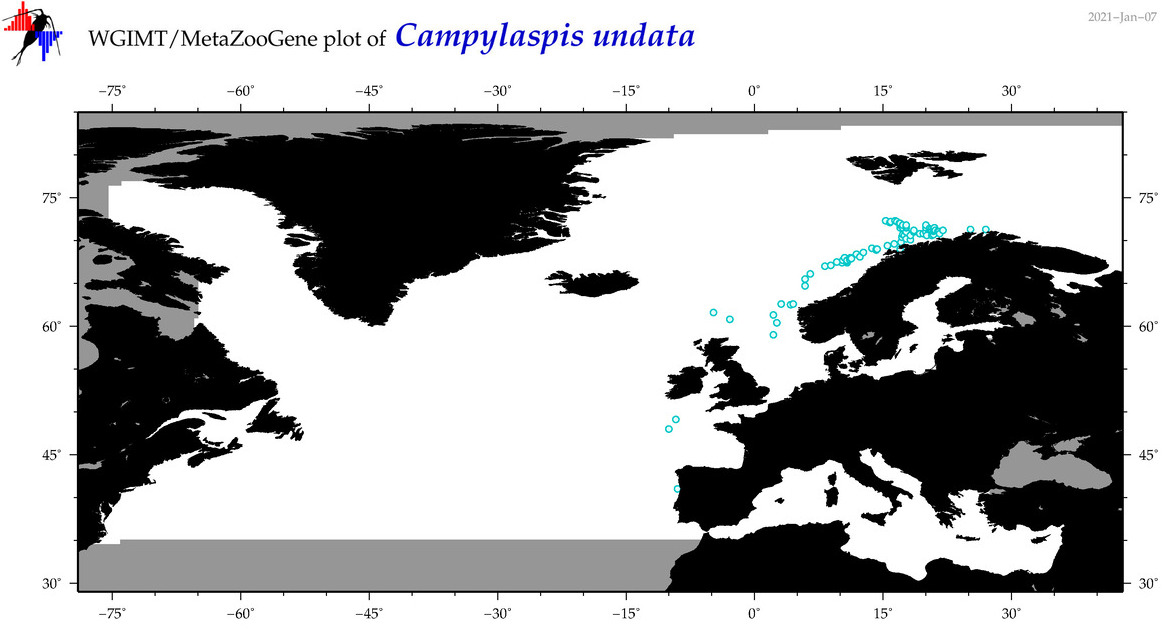
| Campylaspis undata |
Species
(25) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046638
R:1:0:0:0 |
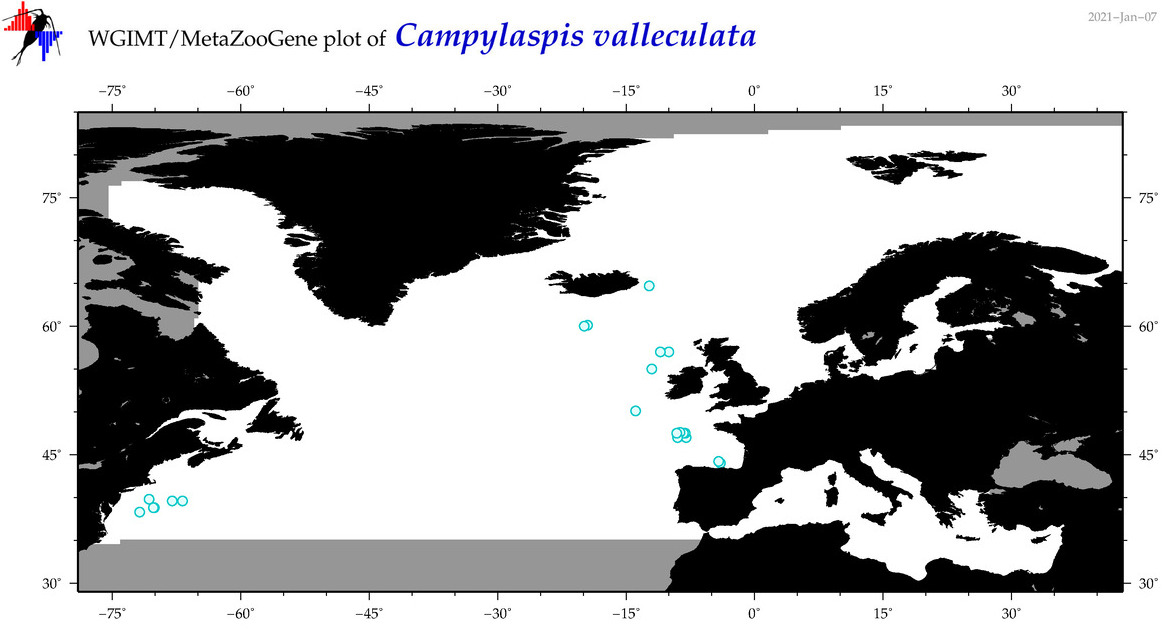
| Campylaspis valleculata |
Species
(26) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046642
R:1:0:0:0 |
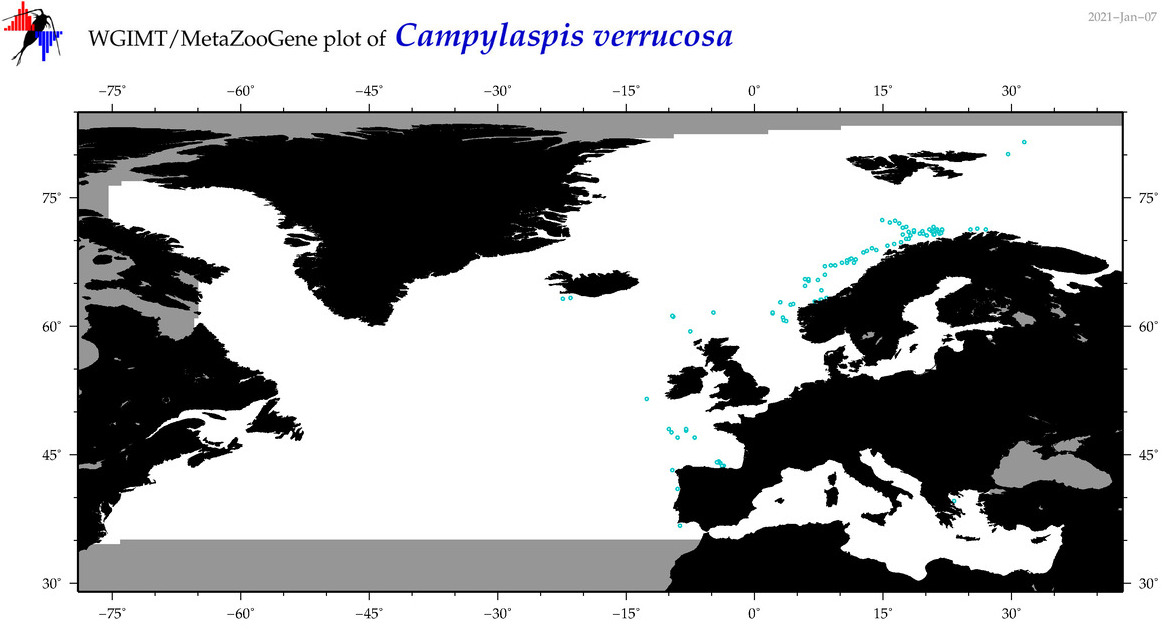
| Campylaspis verrucosa |
Species
(27) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4046643
R:1:0:0:0 |
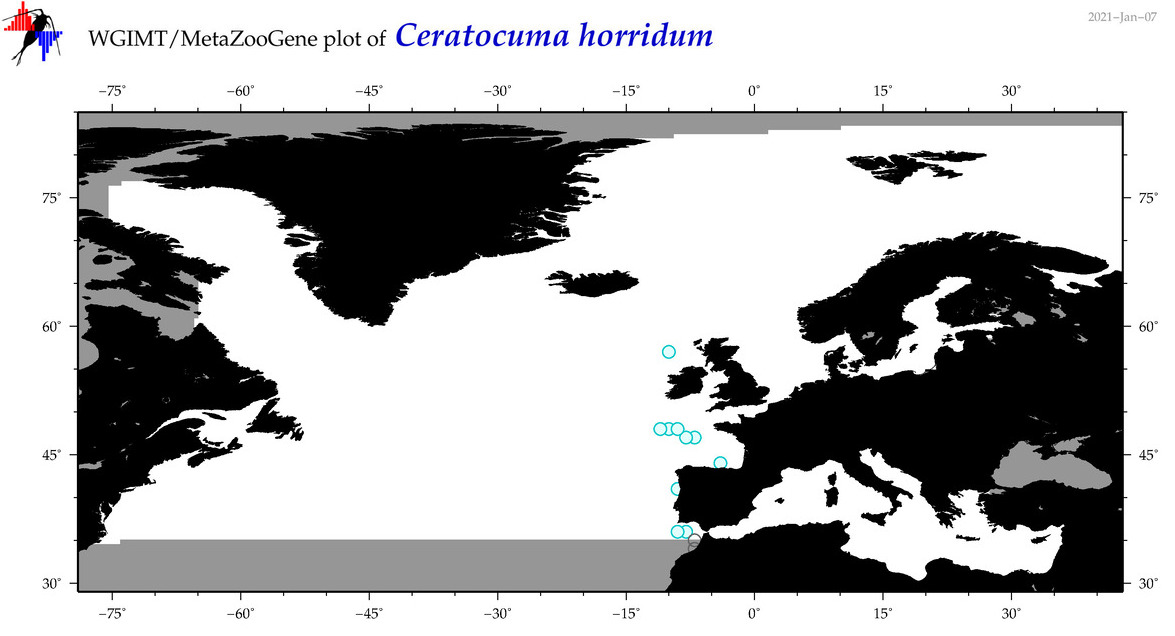
| Ceratocuma horridum |
Species
(28) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4077403
R:1:0:0:0 |
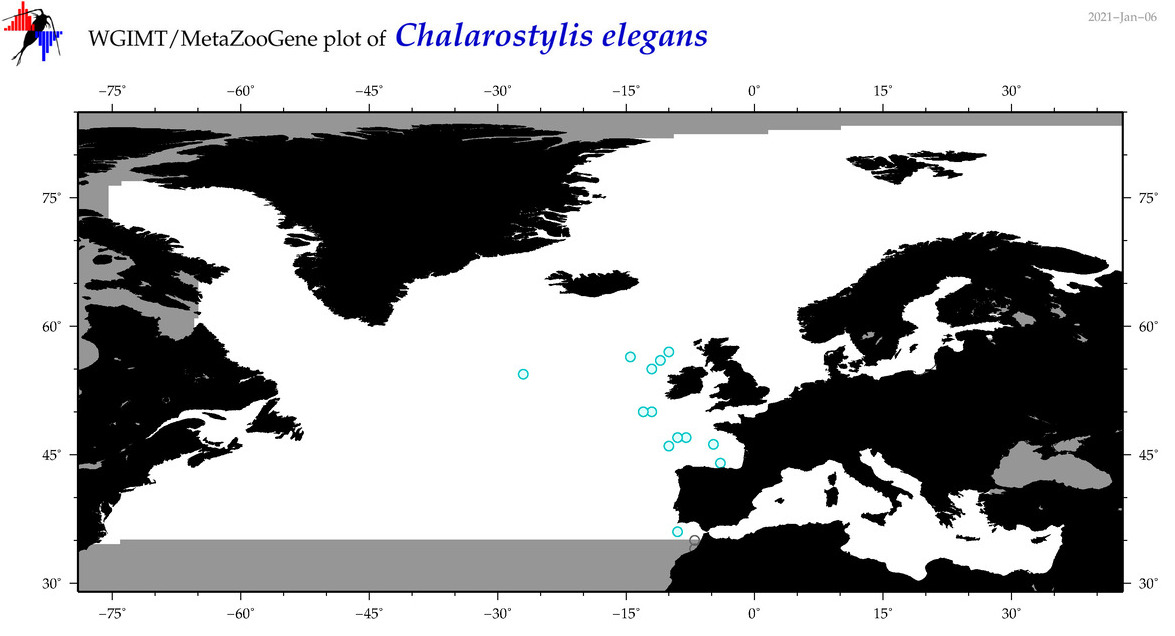
| Chalarostylis elegans |
Species
(29) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4047440
R:1:0:0:0 |
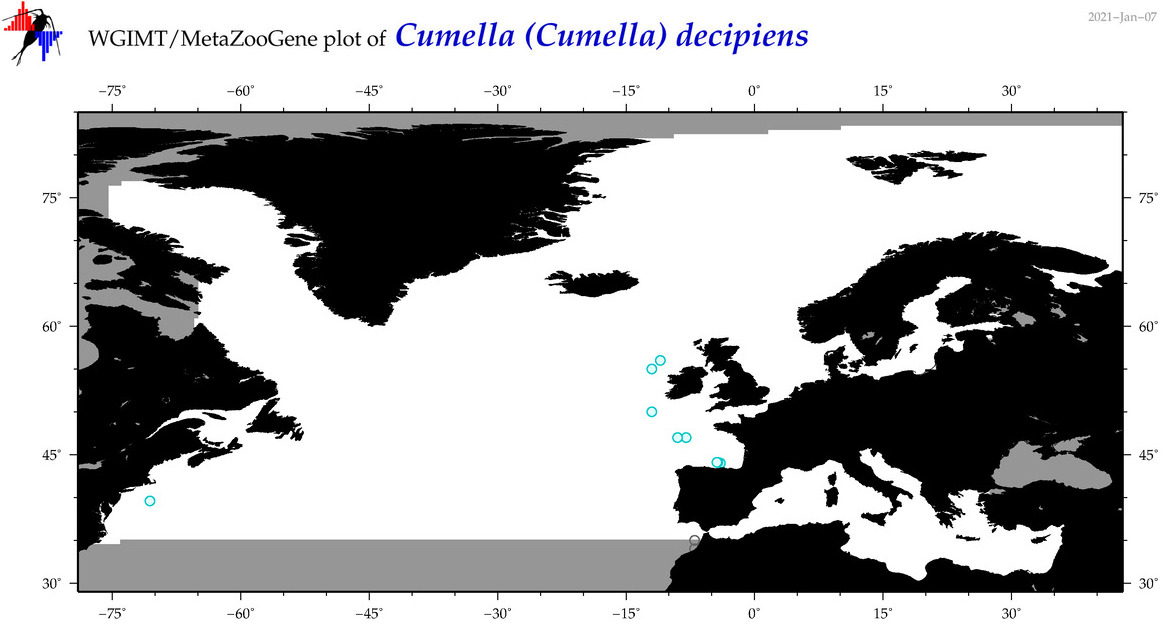
| Cumella (Cumella) decipiens |
Species
(30) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091474
R:1:0:0:0 |
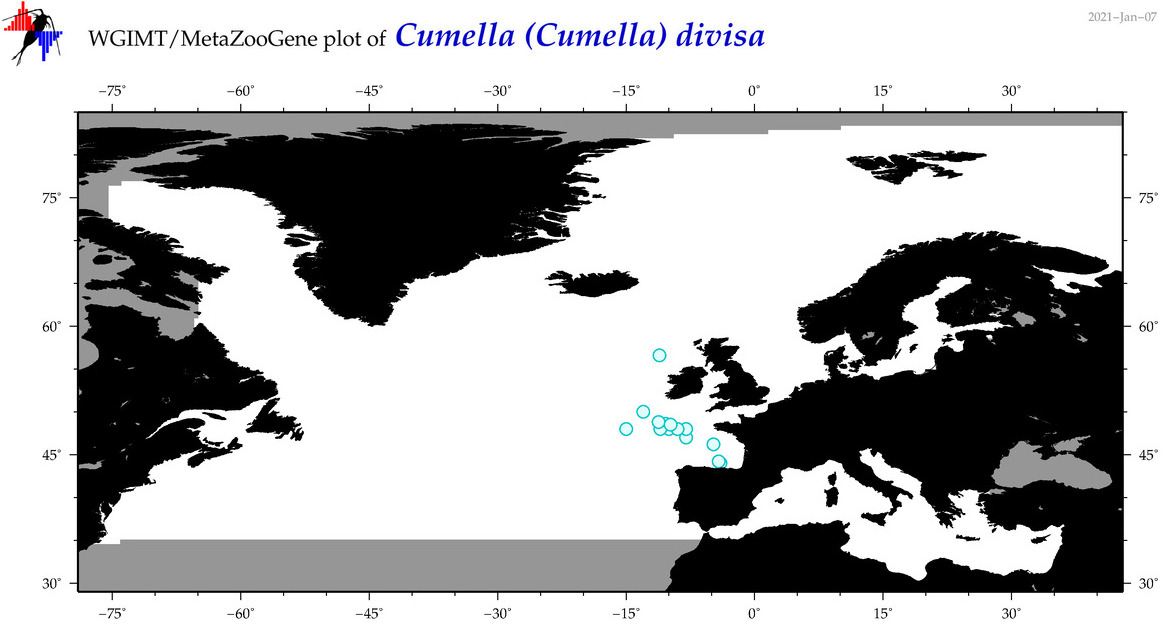
| Cumella (Cumella) divisa |
Species
(31) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094878
R:1:0:0:0 |
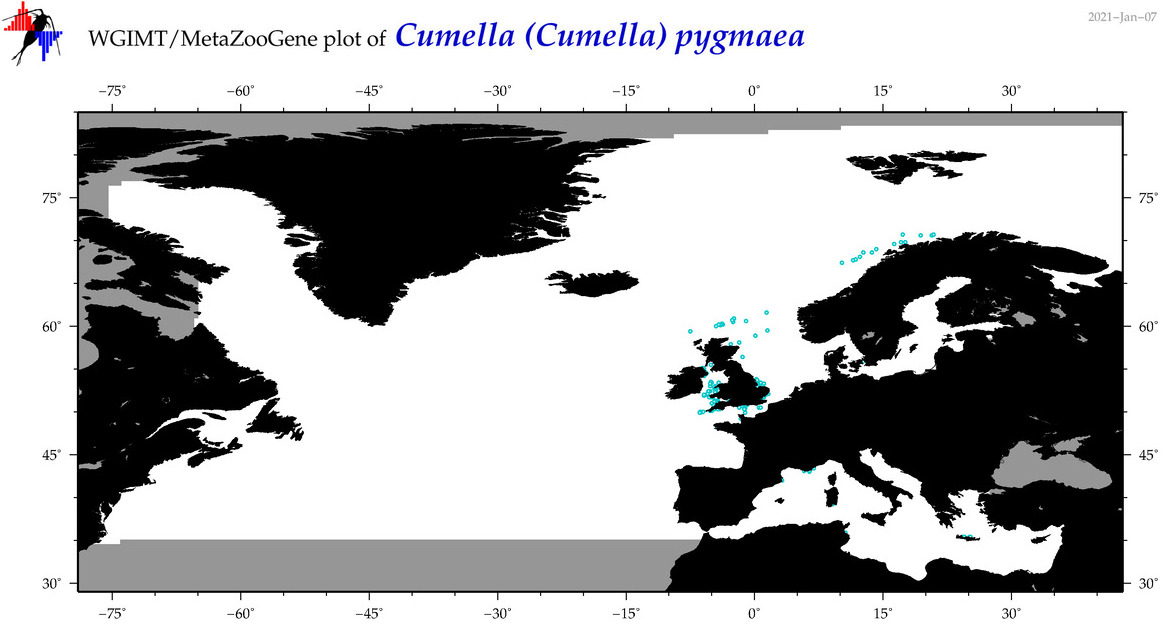
| Cumella (Cumella) pygmaea |
Species
(32) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091476
R:1:0:0:0 |
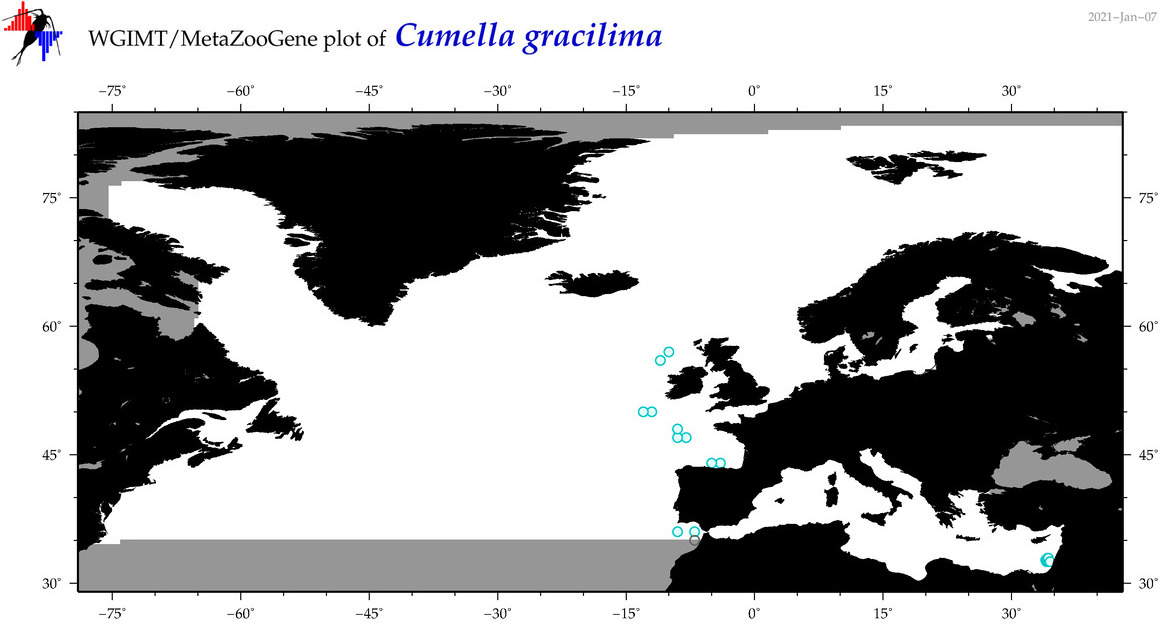
| Cumella gracilima |
Species
(33) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4078025
R:1:0:0:0 |
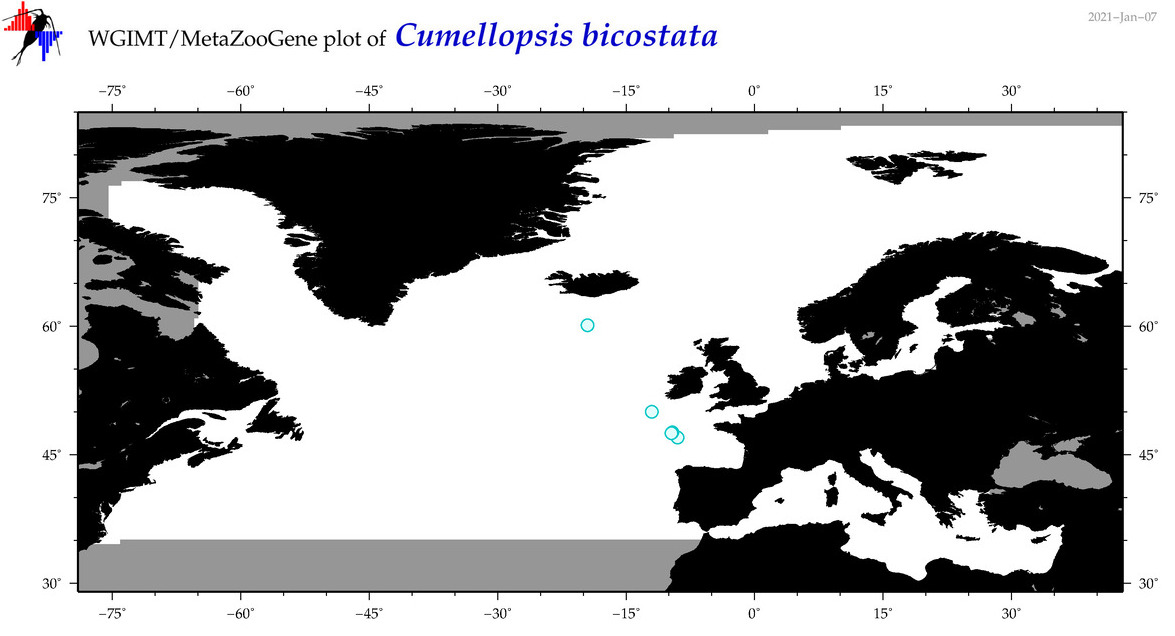
| Cumellopsis bicostata |
Species
(34) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048823
R:1:0:0:0 |
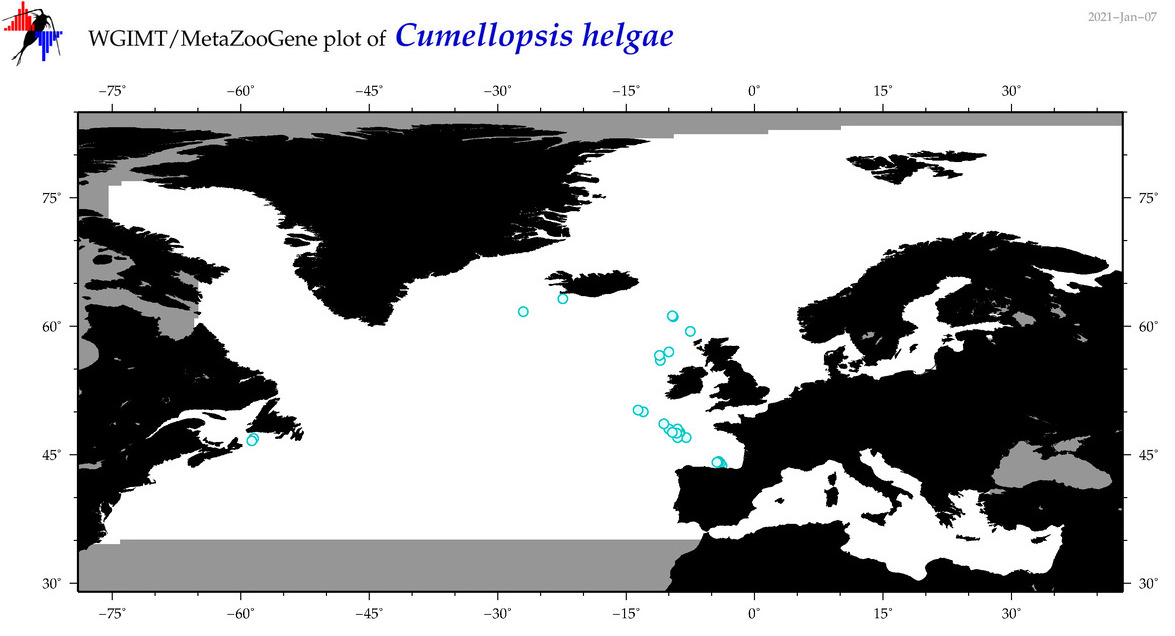
| Cumellopsis helgae |
Species
(35) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048825
R:1:0:0:0 |
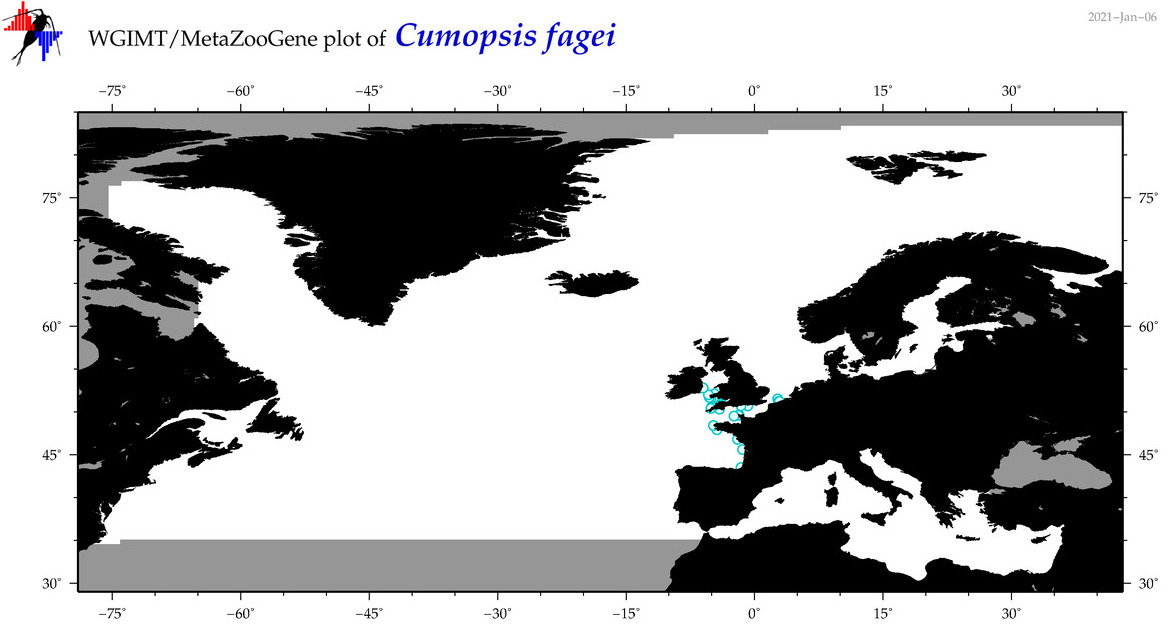
| Cumopsis fagei |
Species
(36) |
COI
12S
16S
18S
28S
|
COI = 0
12S = 0
16S = 1
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048828
R:1:0:0:0 |
| Cumopsis goodsir |
Species
(37) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4096287
R:1:0:0:0 |
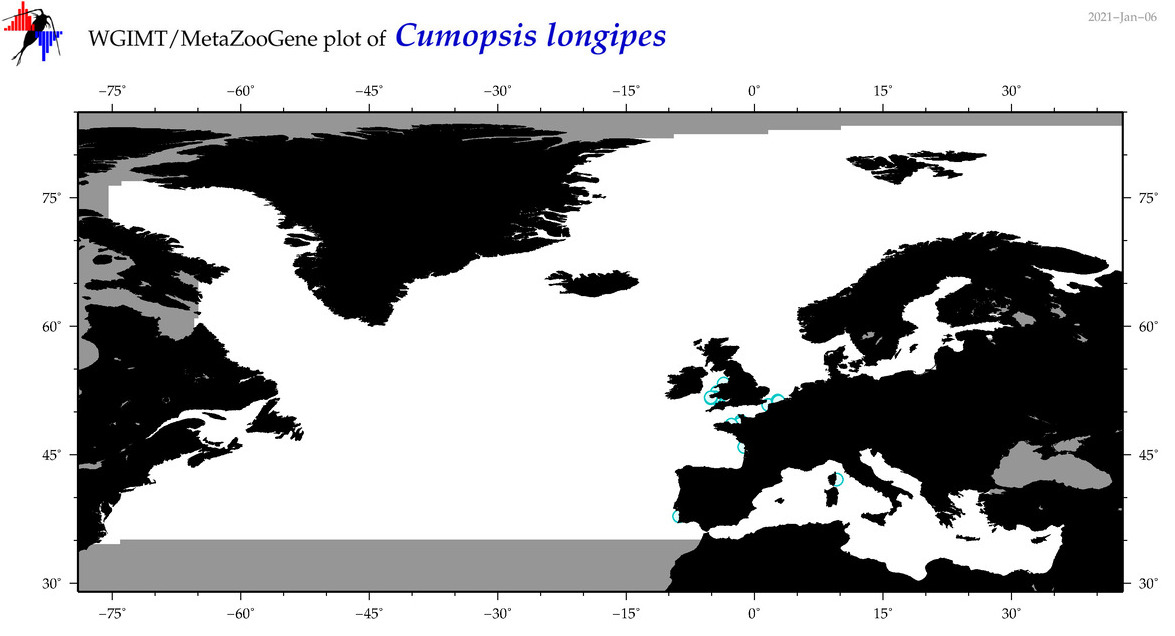
| Cumopsis longipes |
Species
(38) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048829
R:1:0:0:0 |
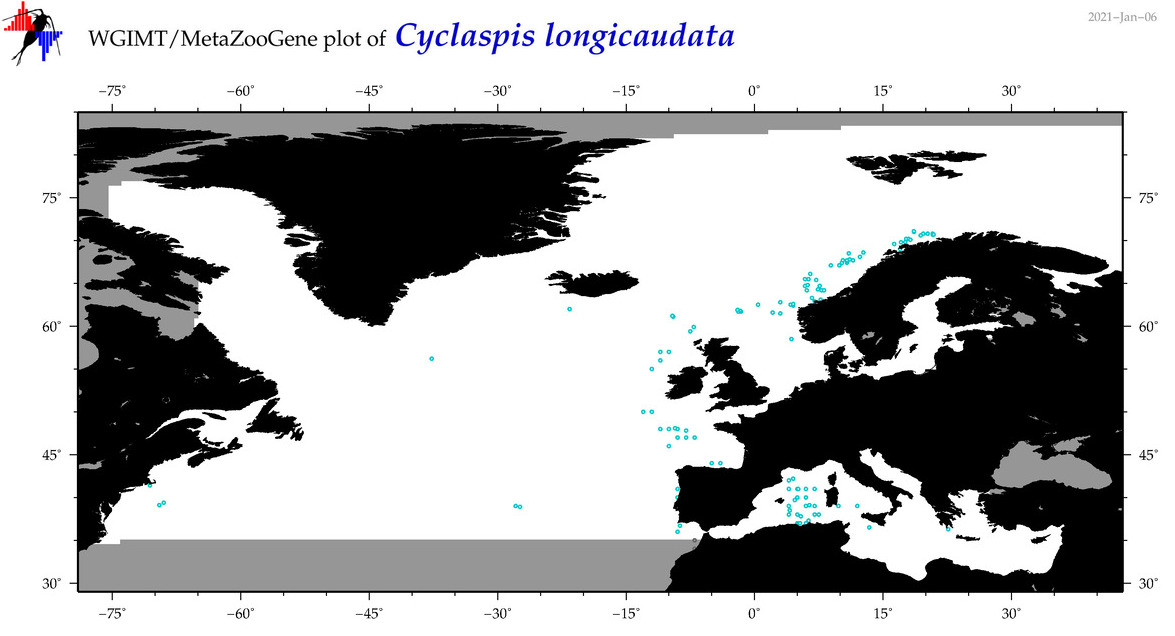
| Cyclaspis longicaudata |
Species
(39) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 2
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048905
R:1:0:0:0 |
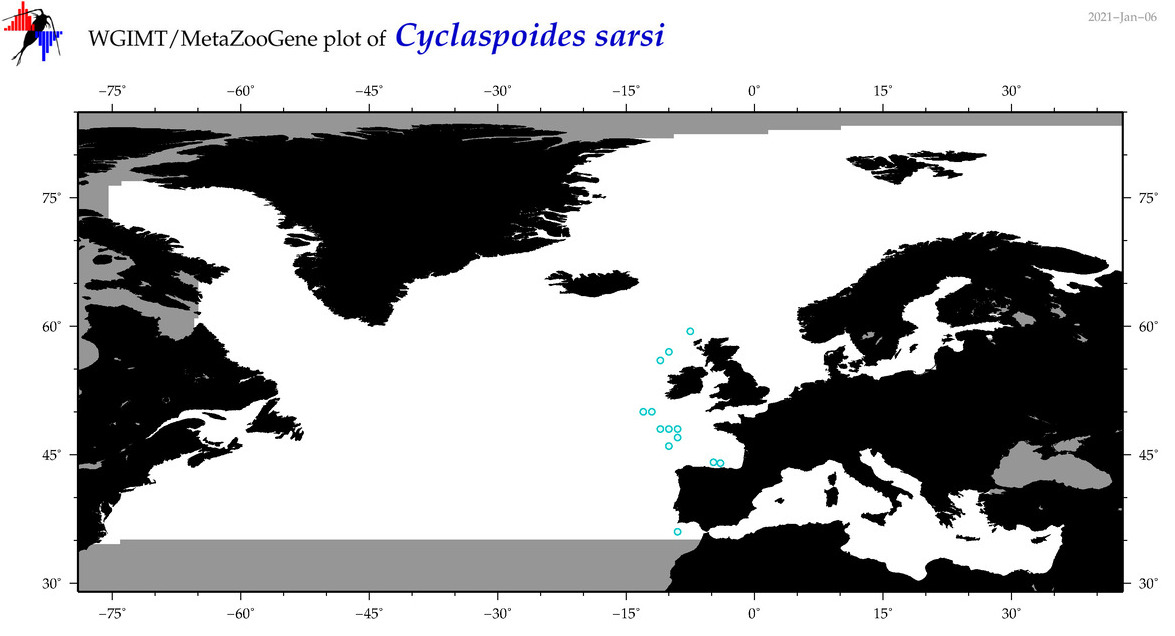
| Cyclaspoides sarsi |
Species
(40) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4048947
R:1:0:0:0 |
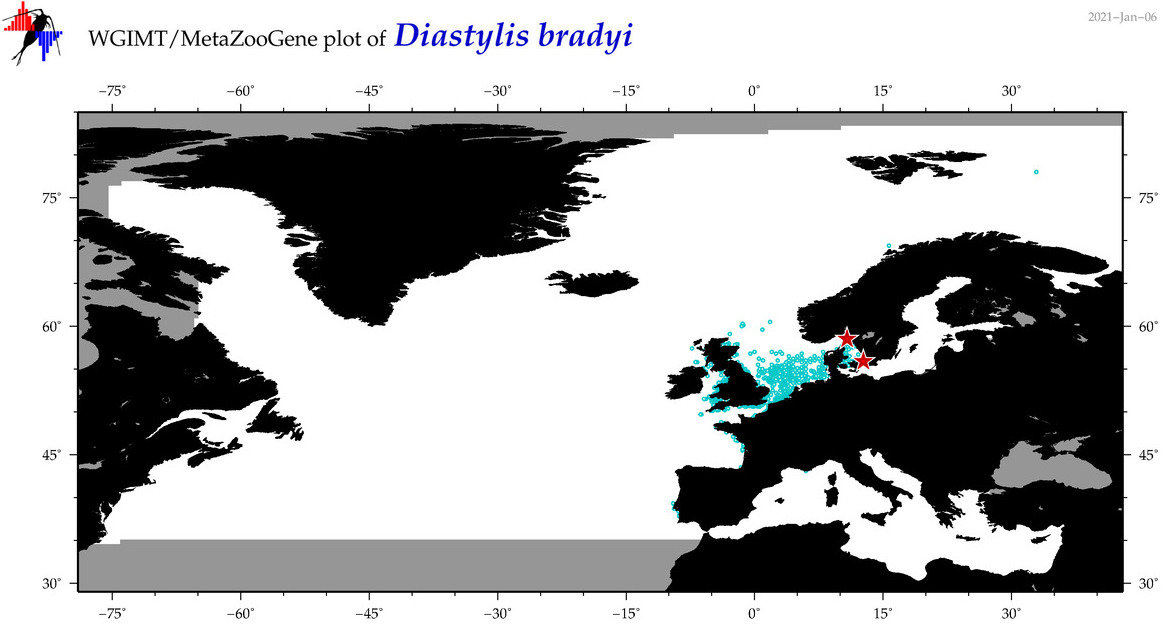
| Diastylis bradyi |
Species
(41) |
COI
12S
16S
18S
28S
|
COI = 9
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4021785
R:1:1:0:0 |
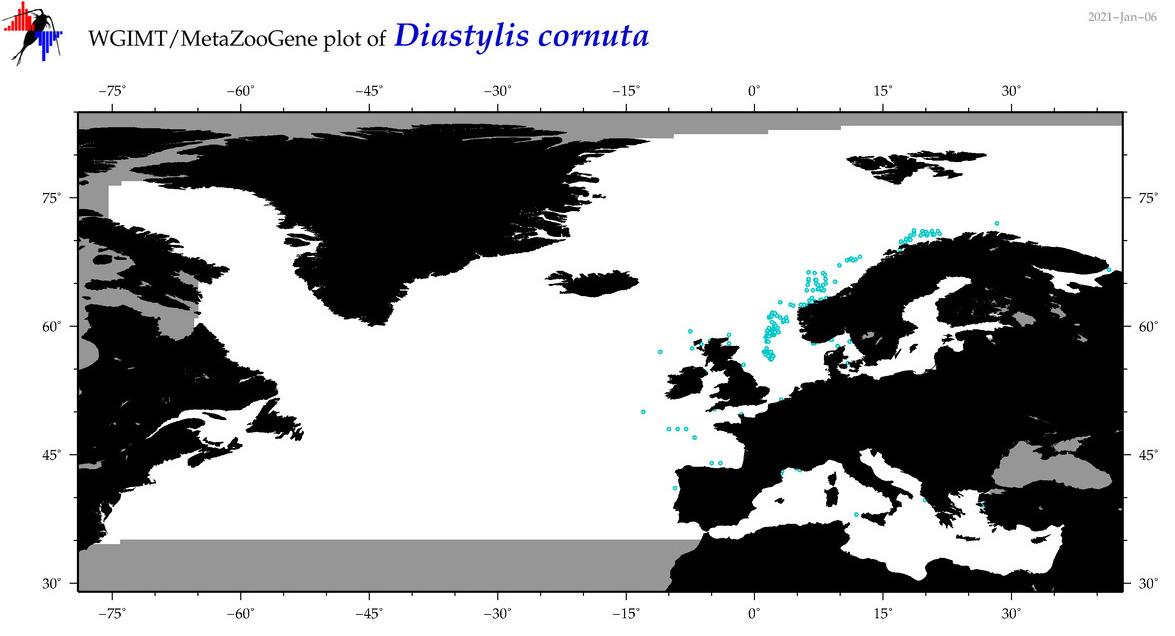
| Diastylis cornuta |
Species
(42) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049580
R:1:1:0:0 |
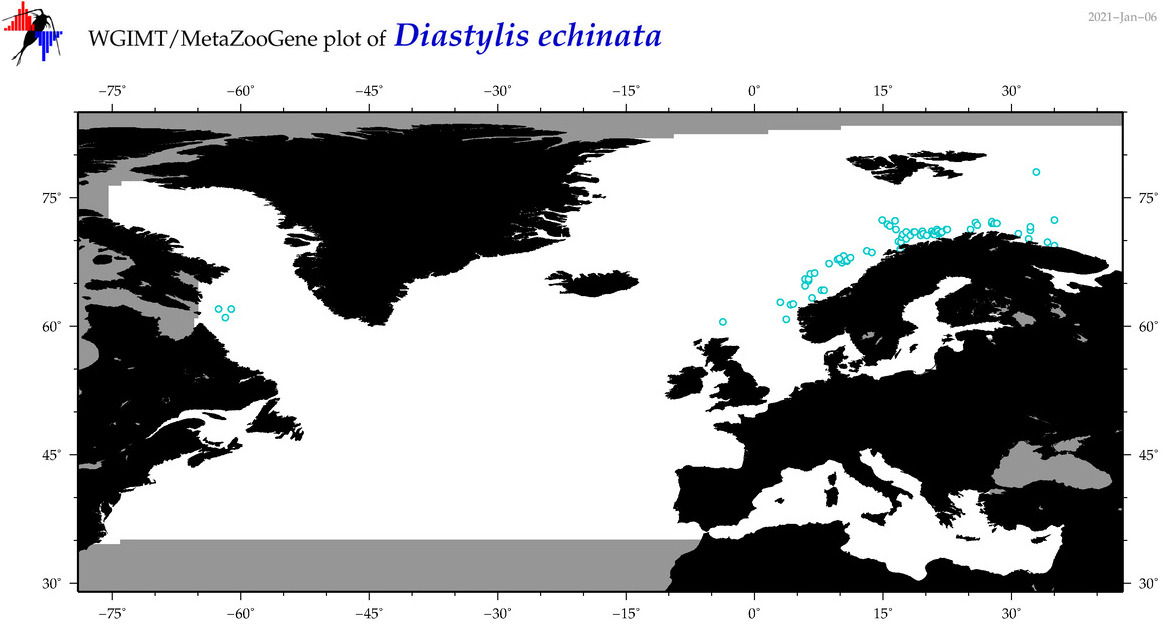
| Diastylis echinata |
Species
(43) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049585
R:1:1:0:0 |
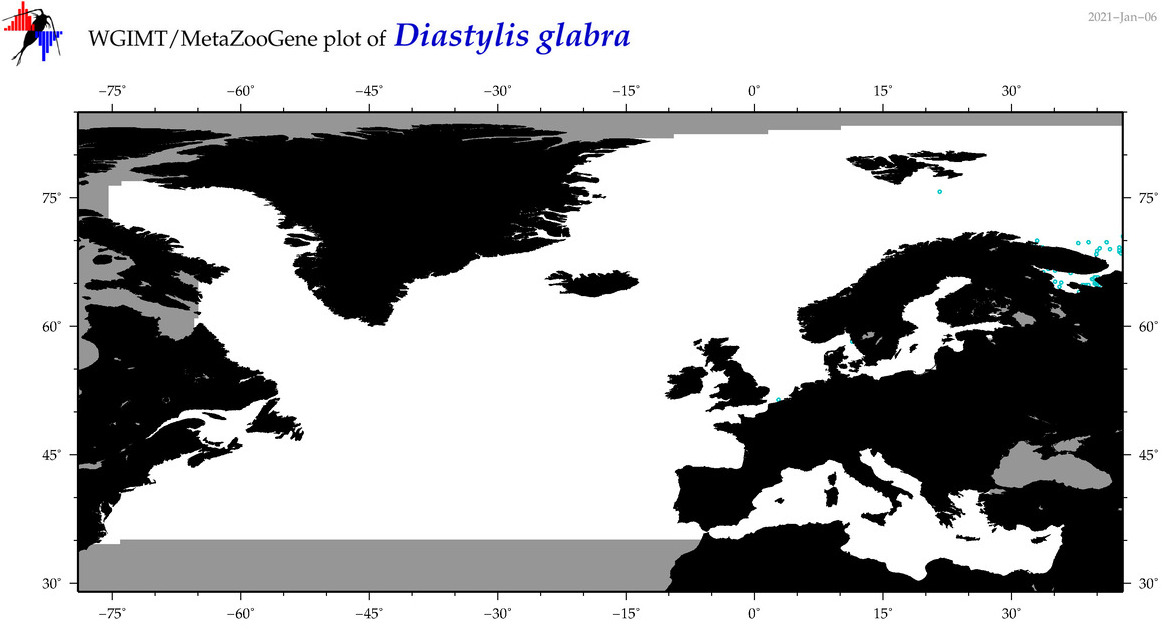
| Diastylis glabra |
Species
(44) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049591
R:1:1:0:0 |
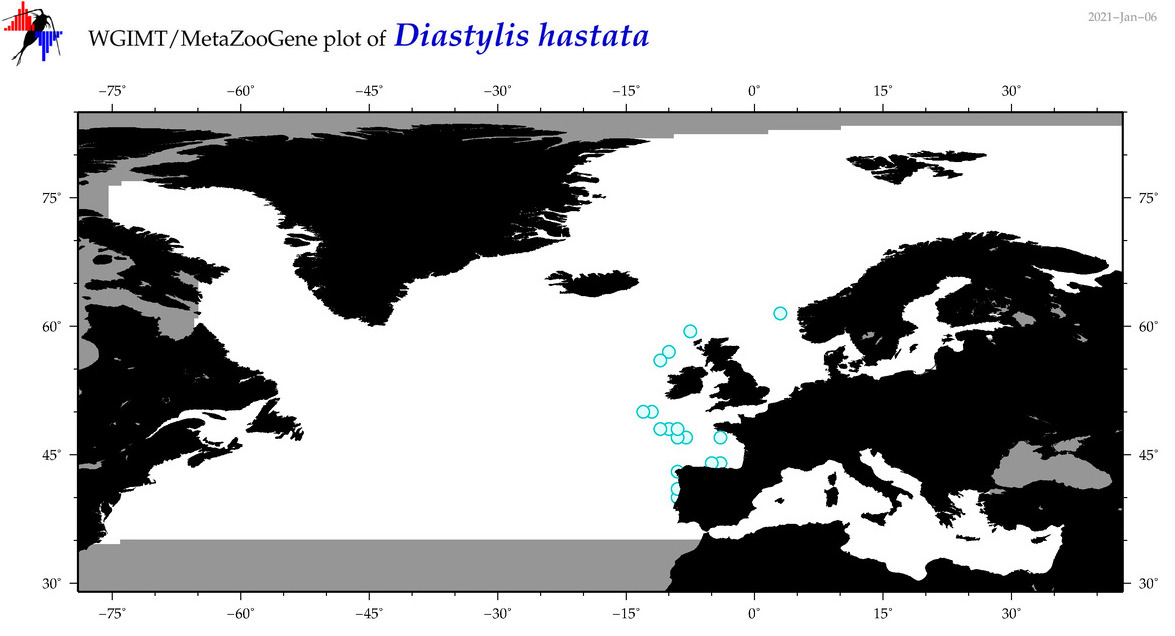
| Diastylis hastata |
Species
(45) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049593
R:1:1:0:0 |
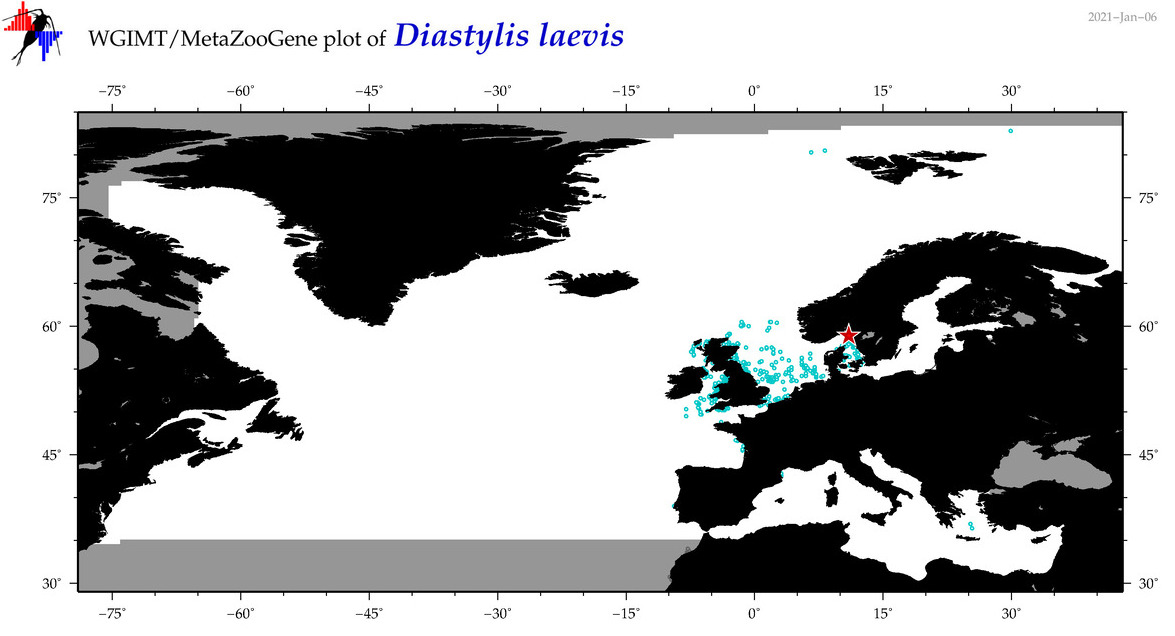
| Diastylis laevis |
Species
(46) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016715
R:1:1:0:0 |
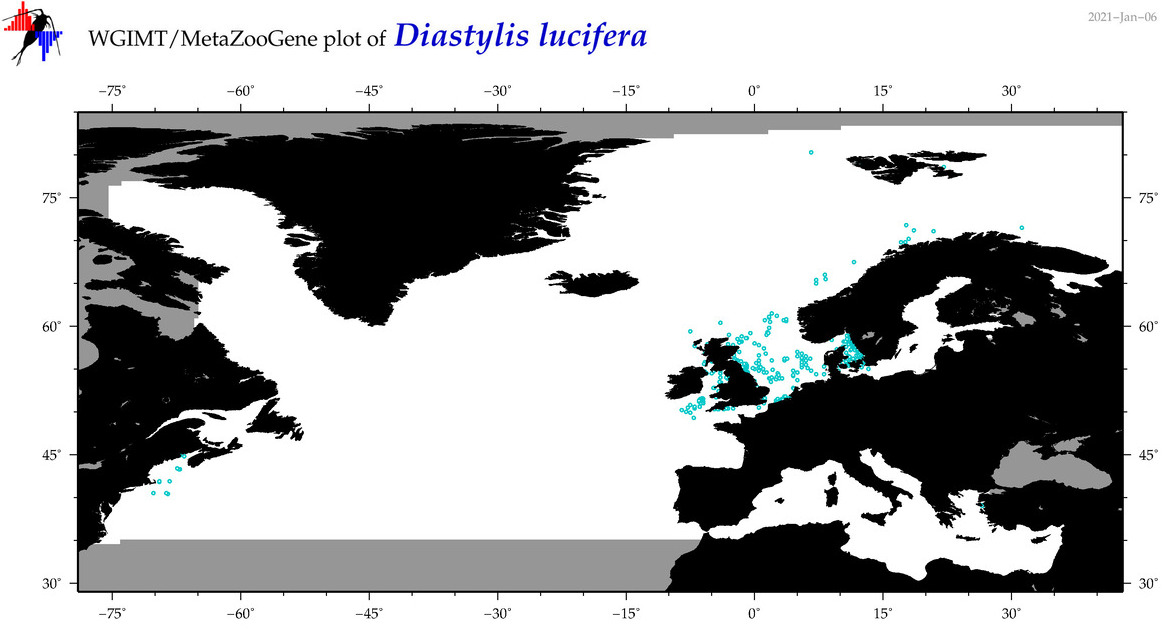
| Diastylis lucifera |
Species
(47) |
COI
12S
16S
18S
28S
|
COI = 10
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016716
R:1:1:0:0 |
| Diastylis rathkei |
Species
(48) |
COI
12S
16S
18S
28S
|
COI = 10
12S = 0
16S = 1
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4011179
R:1:1:0:0 |
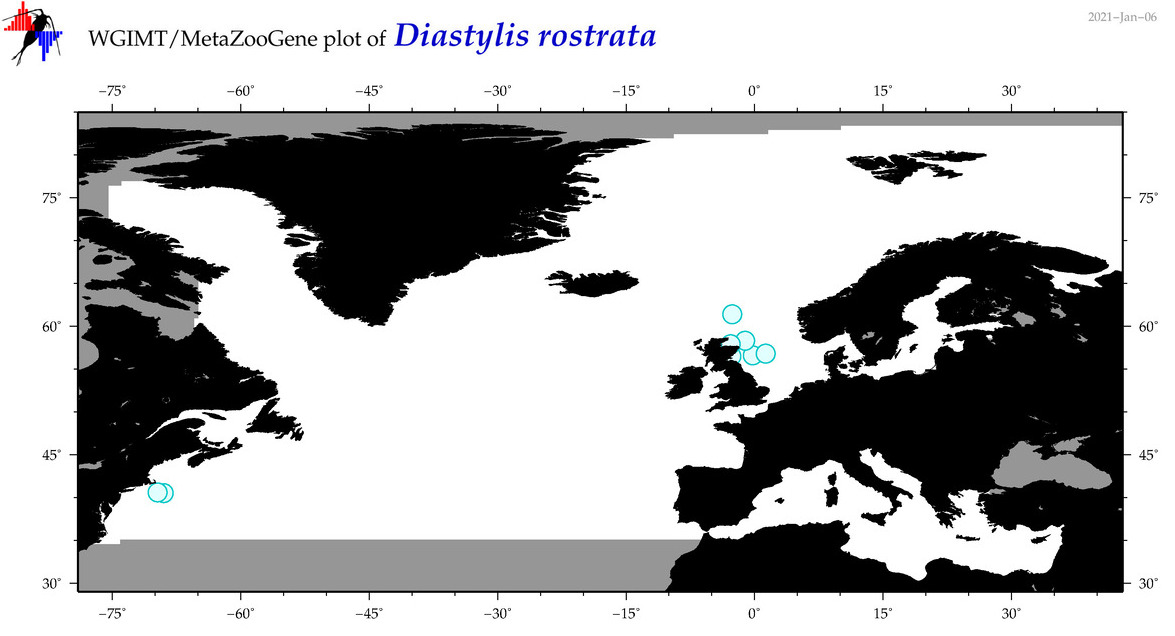
| Diastylis rostrata |
Species
(49) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049620
R:1:1:0:0 |
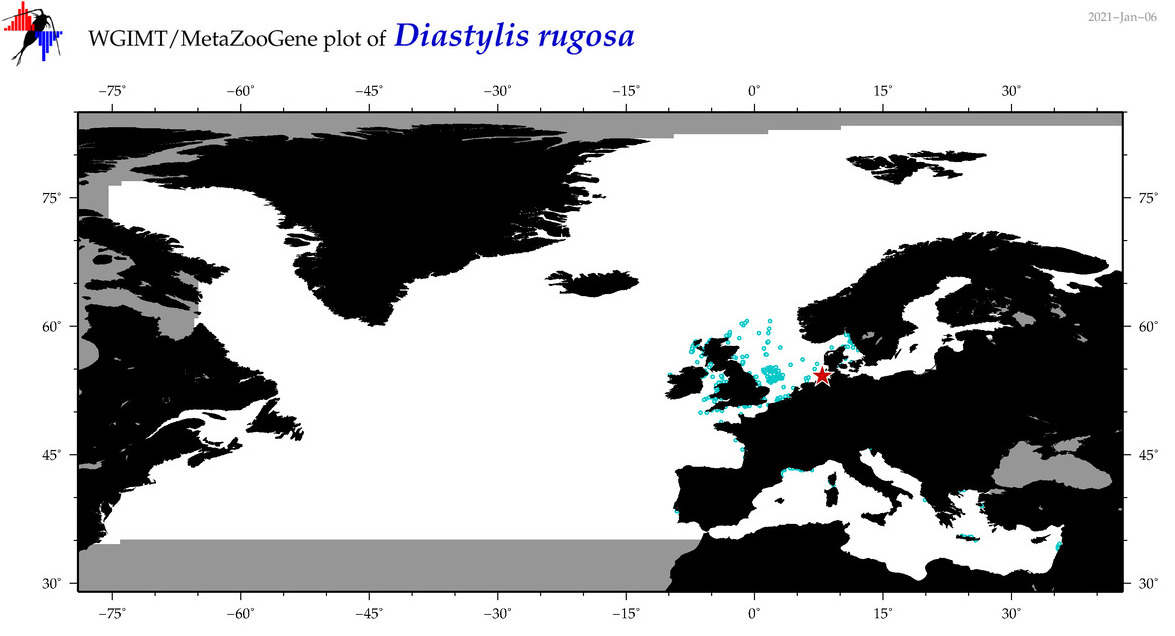
| Diastylis rugosa |
Species
(50) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4011180
R:1:1:0:0 |
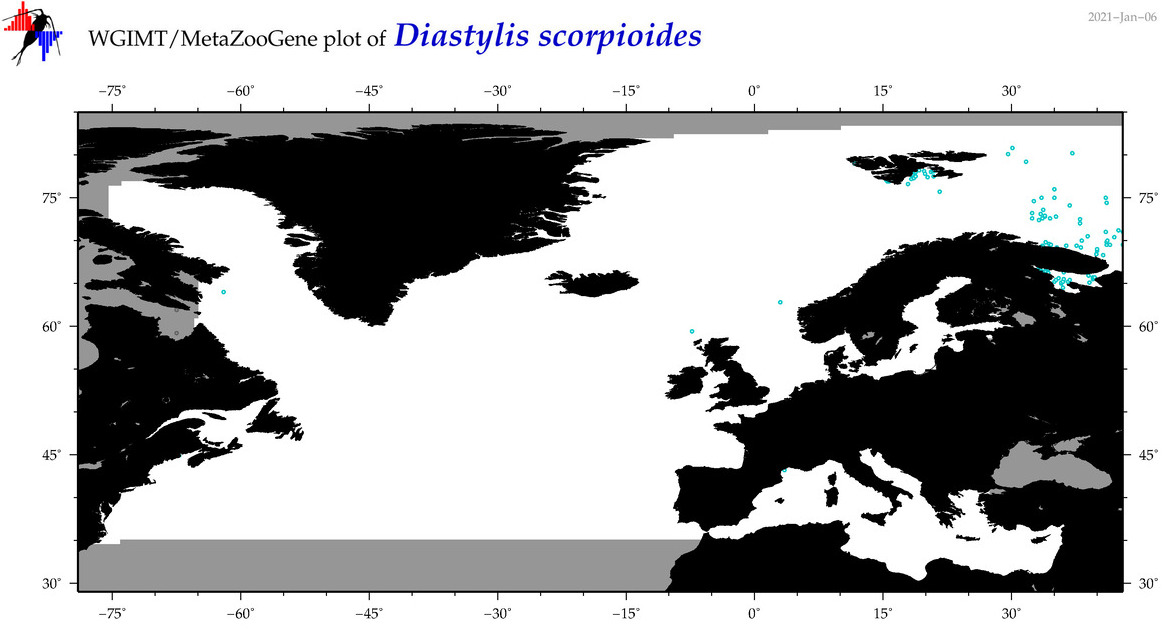
| Diastylis scorpioides |
Species
(51) |
COI
12S
16S
18S
28S
|
COI = 5
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049622
R:1:1:0:0 |
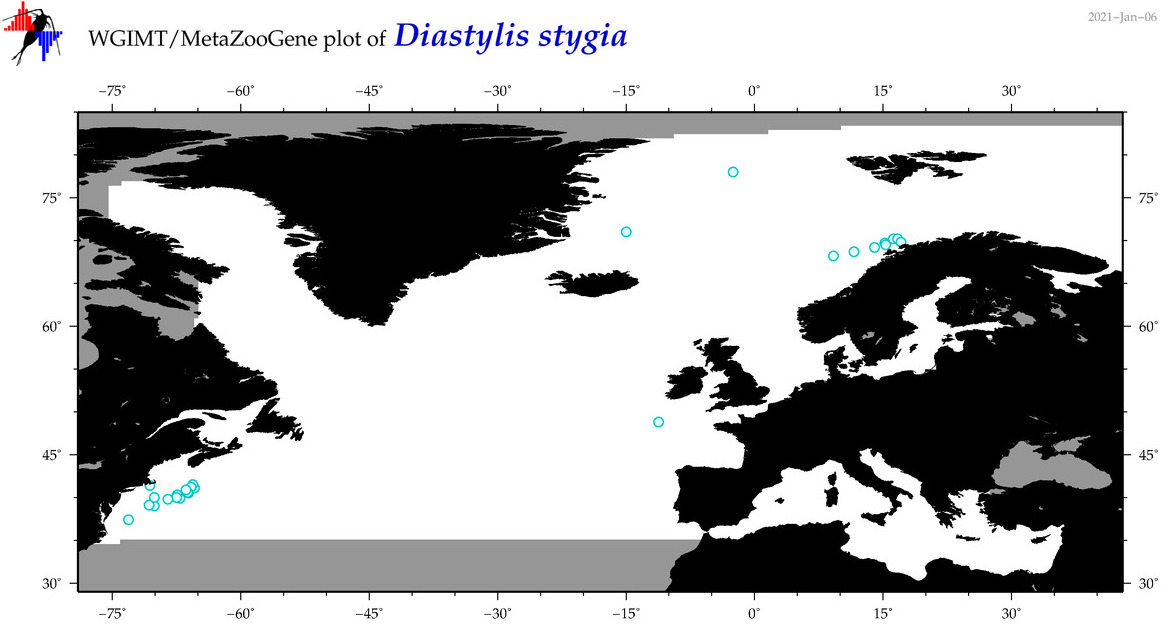
| Diastylis stygia |
Species
(52) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049626
R:1:1:0:0 |
| Diastylis tenuicauda |
Species
(53) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4049630
R:1:1:0:0 |
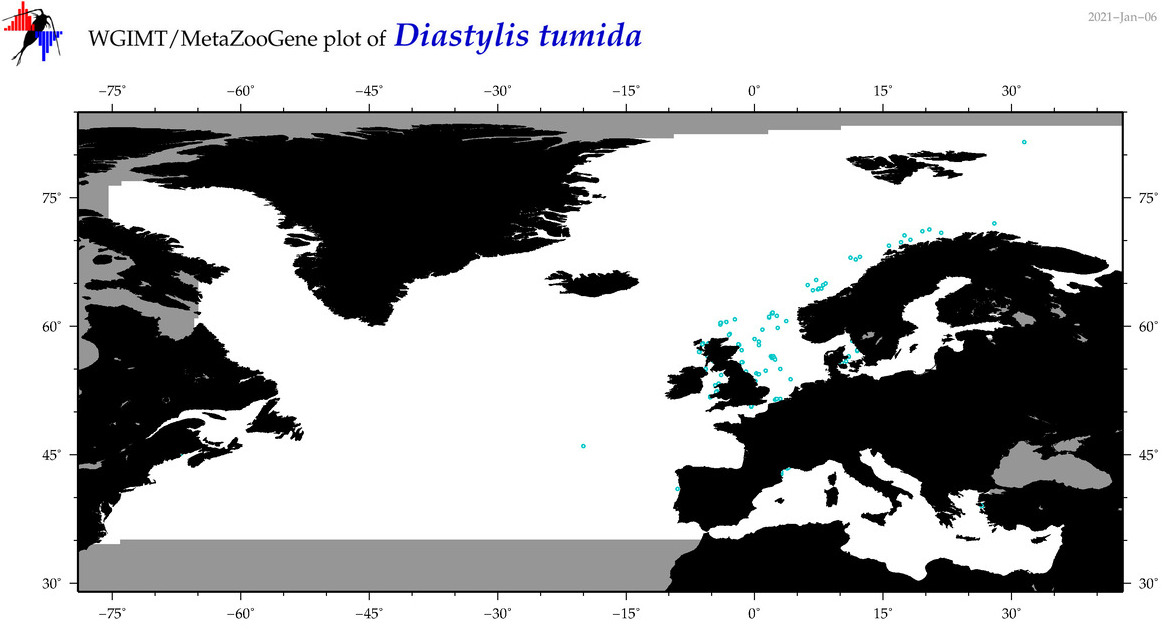
| Diastylis tumida |
Species
(54) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016719
R:1:1:0:0 |
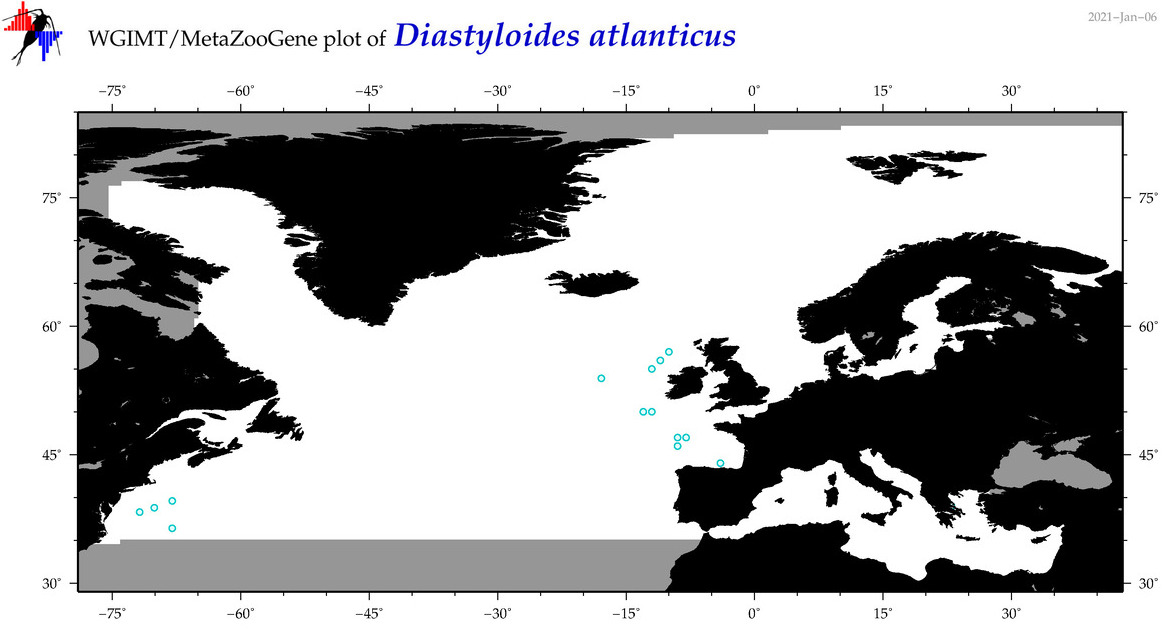
| Diastyloides atlanticus |
Species
(55) |
COI
12S
16S
18S
28S
|
COI = 0
12S = 0
16S = 2
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049634
R:1:1:0:0 |
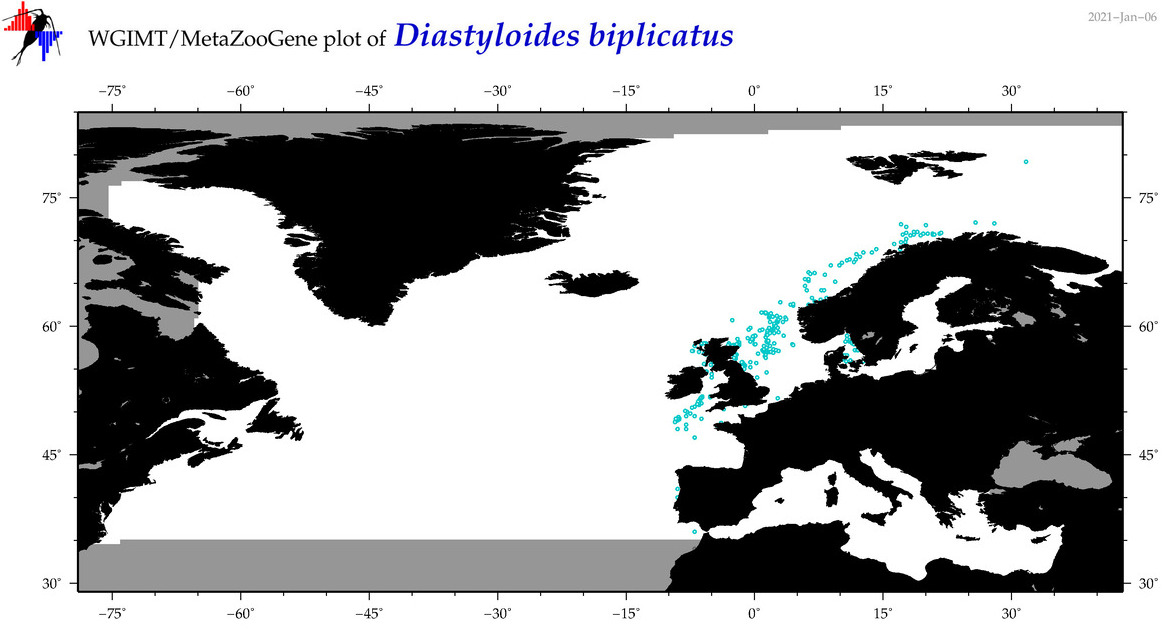
| Diastyloides biplicatus |
Species
(56) |
COI
12S
16S
18S
28S
|
COI = 7
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016720
R:1:1:0:0 |
| Diastyloides scaber |
Species
(57) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4049639
R:1:1:0:0 |
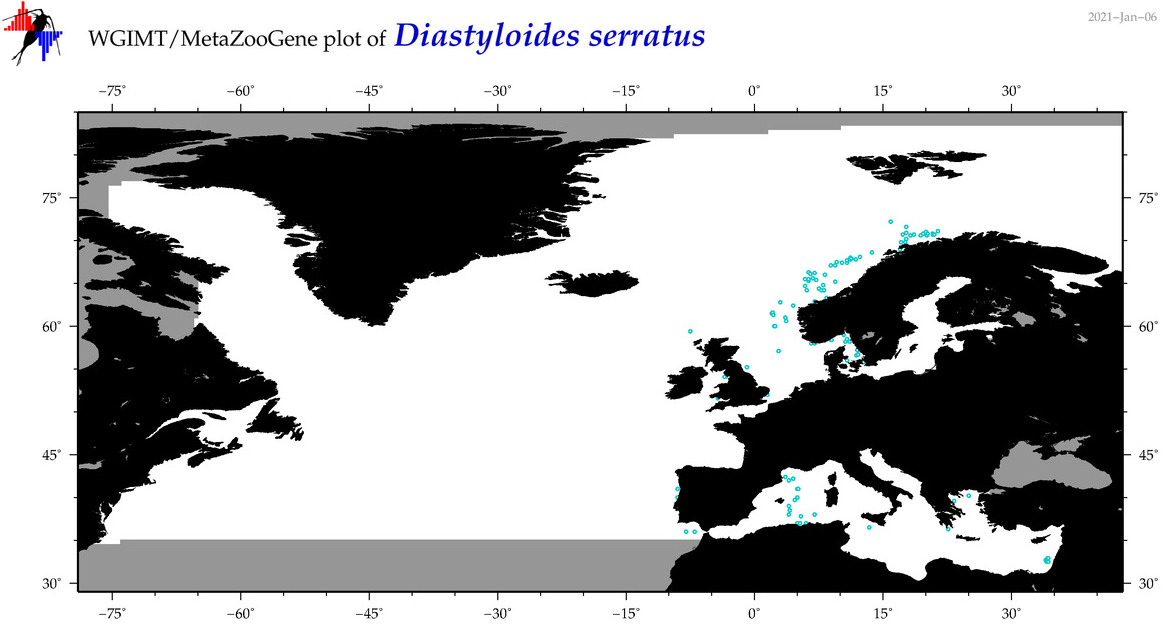
| Diastyloides serratus |
Species
(58) |
COI
12S
16S
18S
28S
|
COI = 7
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016721
R:1:1:0:0 |
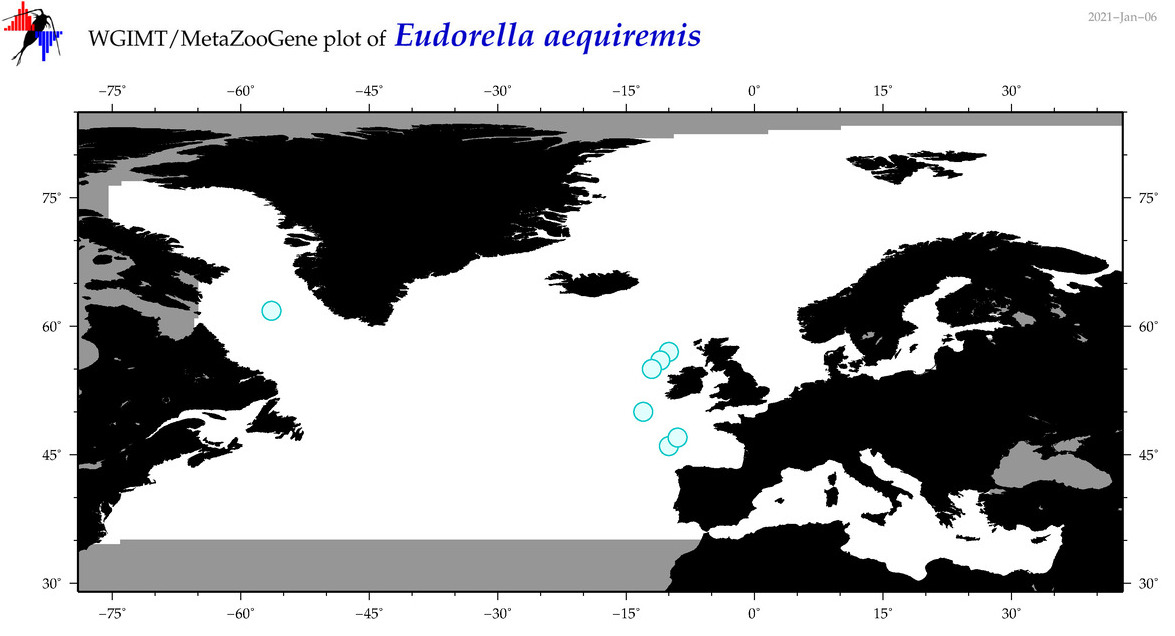
| Eudorella aequiremis |
Species
(59) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4051385
R:1:1:0:0 |
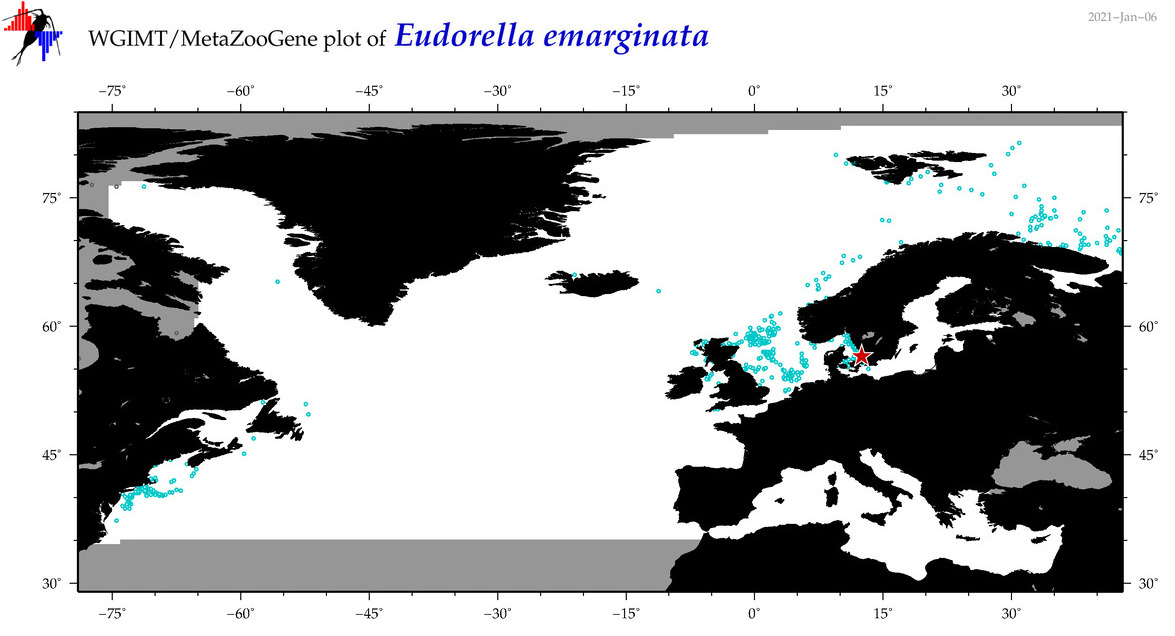
| Eudorella emarginata |
Species
(60) |
COI
12S
16S
18S
28S
|
COI = 162
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4016749
R:1:1:0:0 |
| Eudorella hirsuta |
Species
(61) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
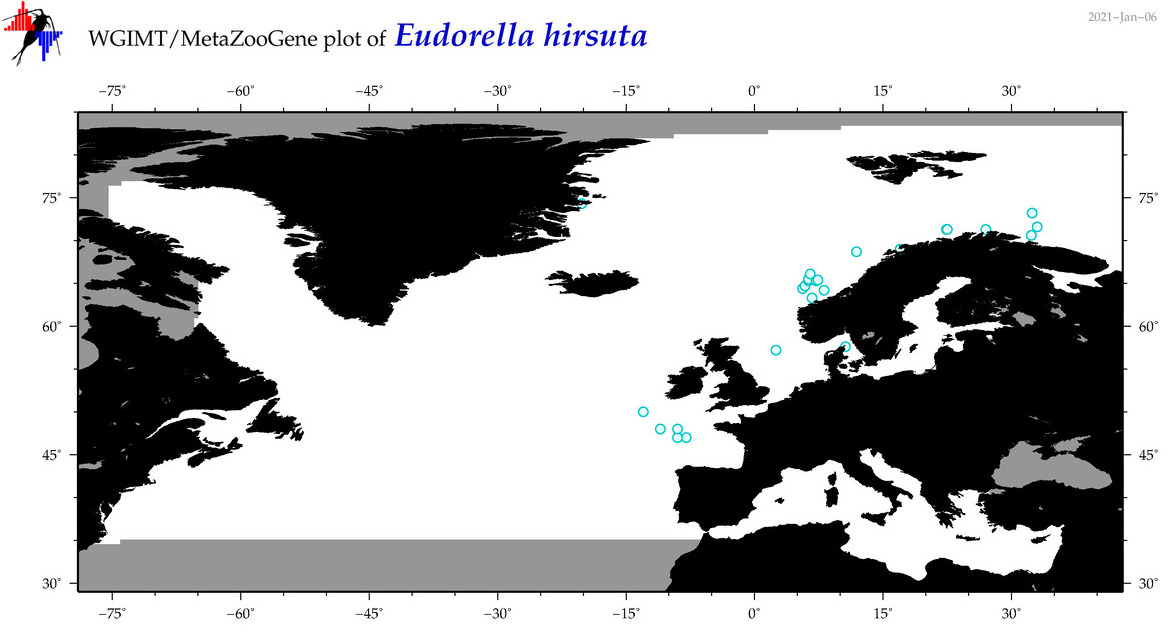 |
accepted
T4051391
R:1:1:0:0 |
| Eudorella intermedia |
Species
(62) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
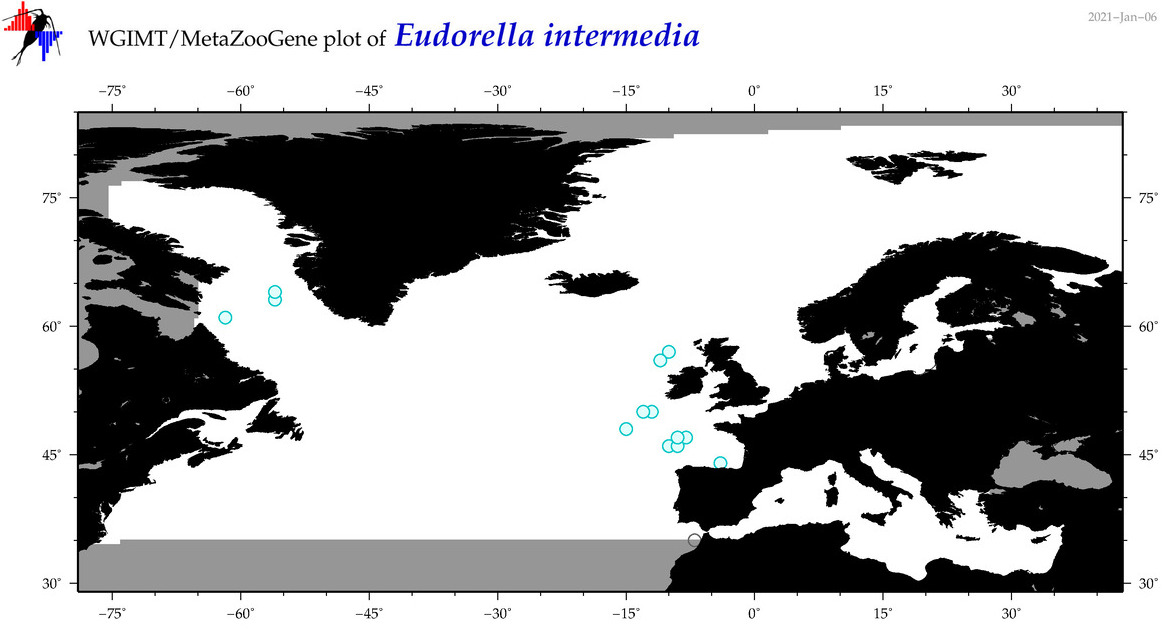 |
accepted
T4051395
R:1:1:0:0 |
| Eudorella parvula |
Species
(63) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
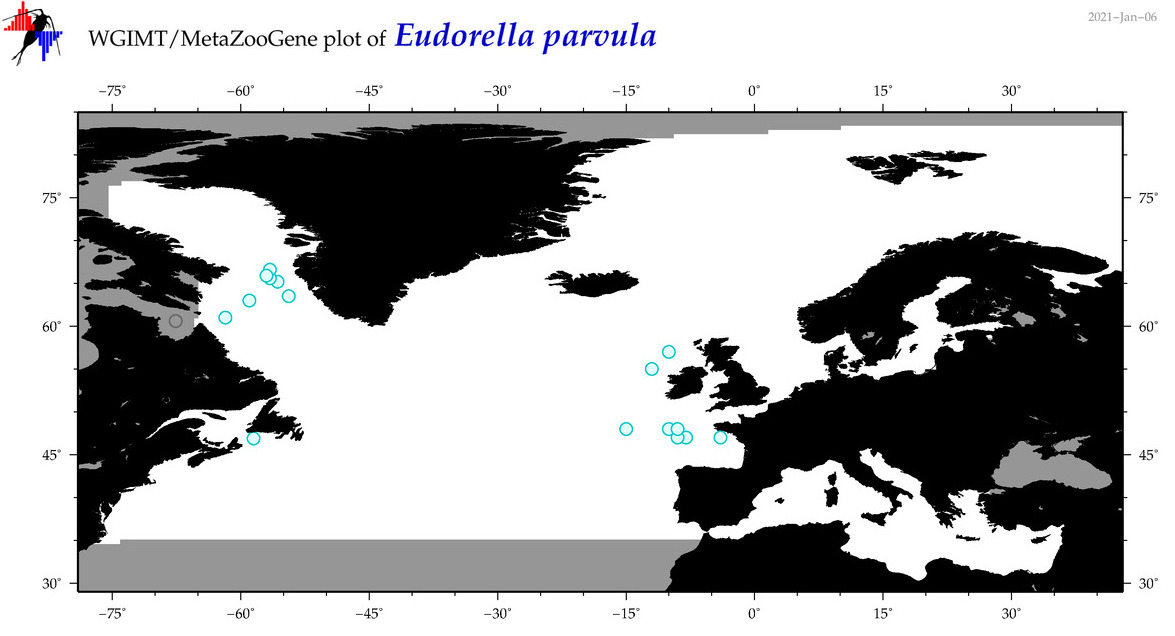 |
accepted
T4051398
R:1:1:0:0 |
| Eudorella truncatula |
Species
(64) |
COI
12S
16S
18S
28S
|
COI = 14
12S = 0
16S = 0
18S = 1
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
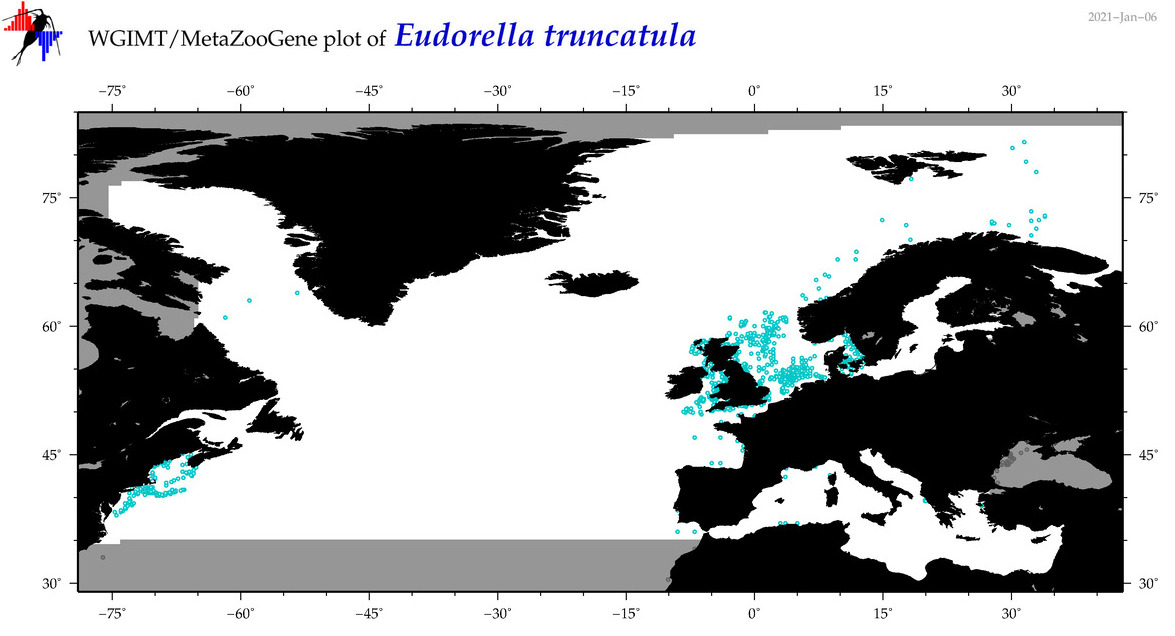 |
accepted
T4051404
R:1:1:0:0 |
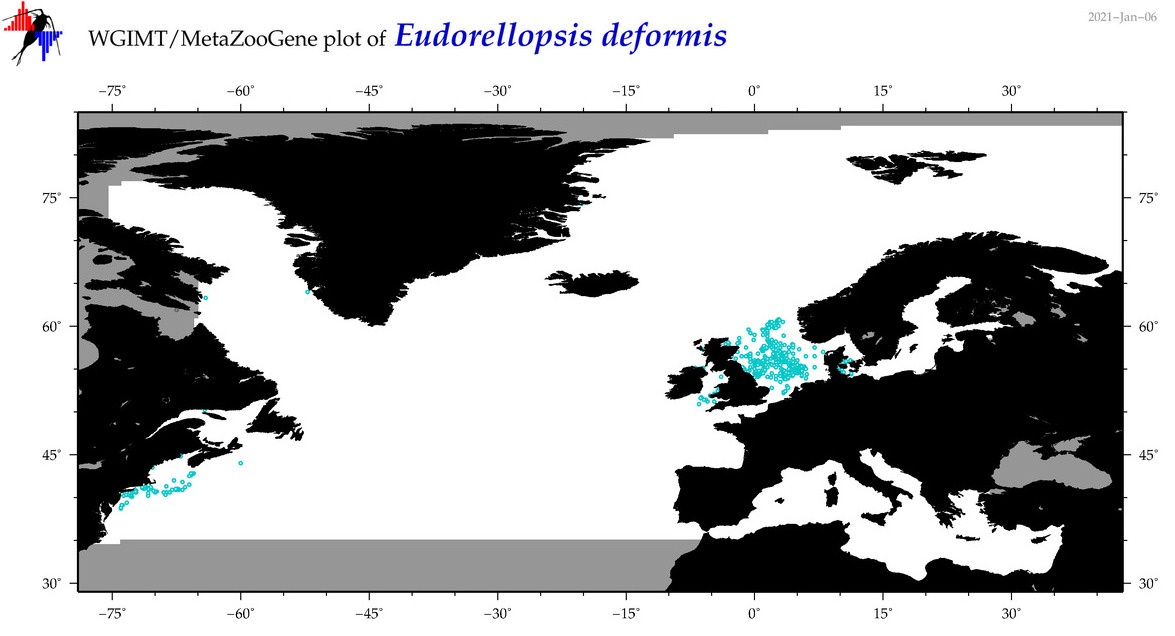
| Eudorellopsis deformis |
Species
(65) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4051406
R:1:1:0:0 |
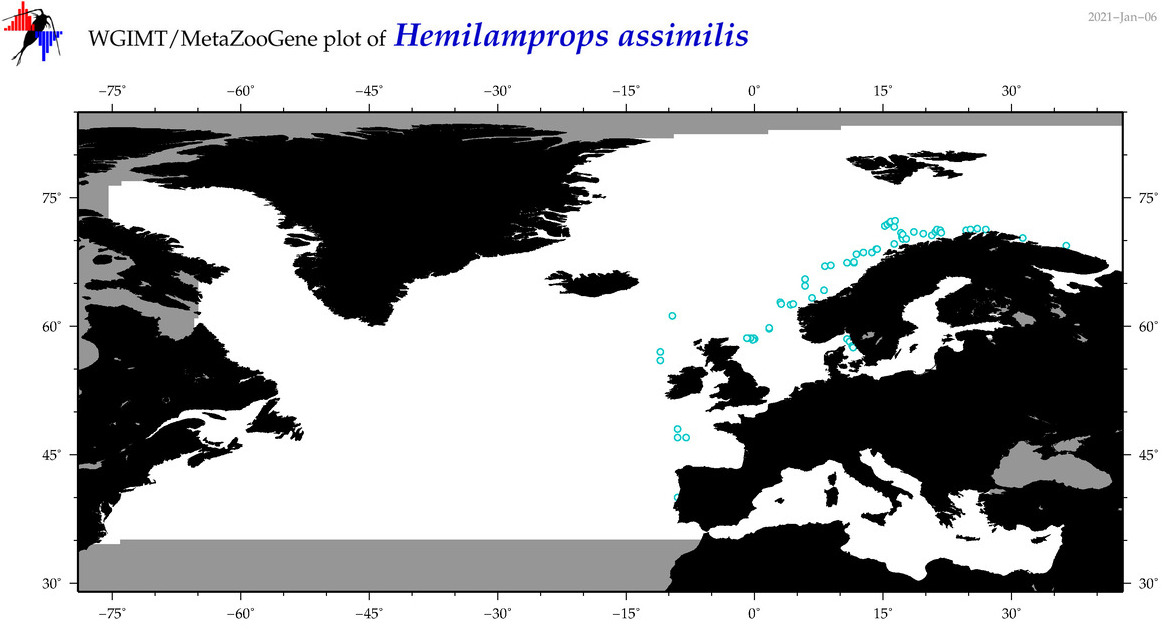
| Hemilamprops assimilis |
Species
(66) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4053841
R:1:0:0:0 |
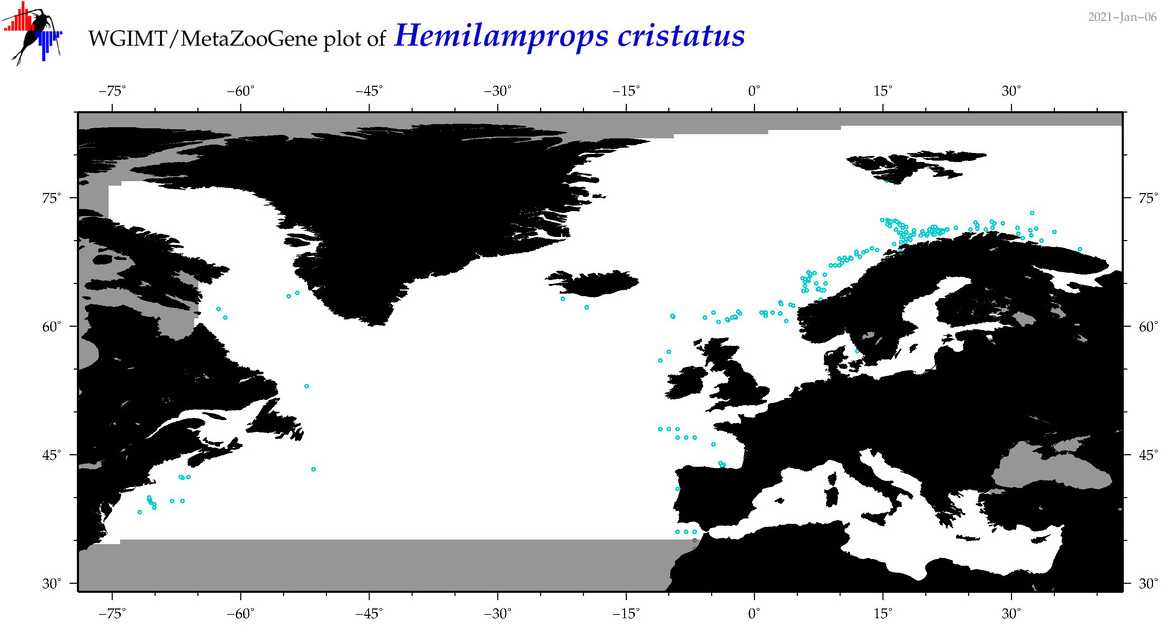
| Hemilamprops cristatus |
Species
(67) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4053844
R:1:0:0:0 |
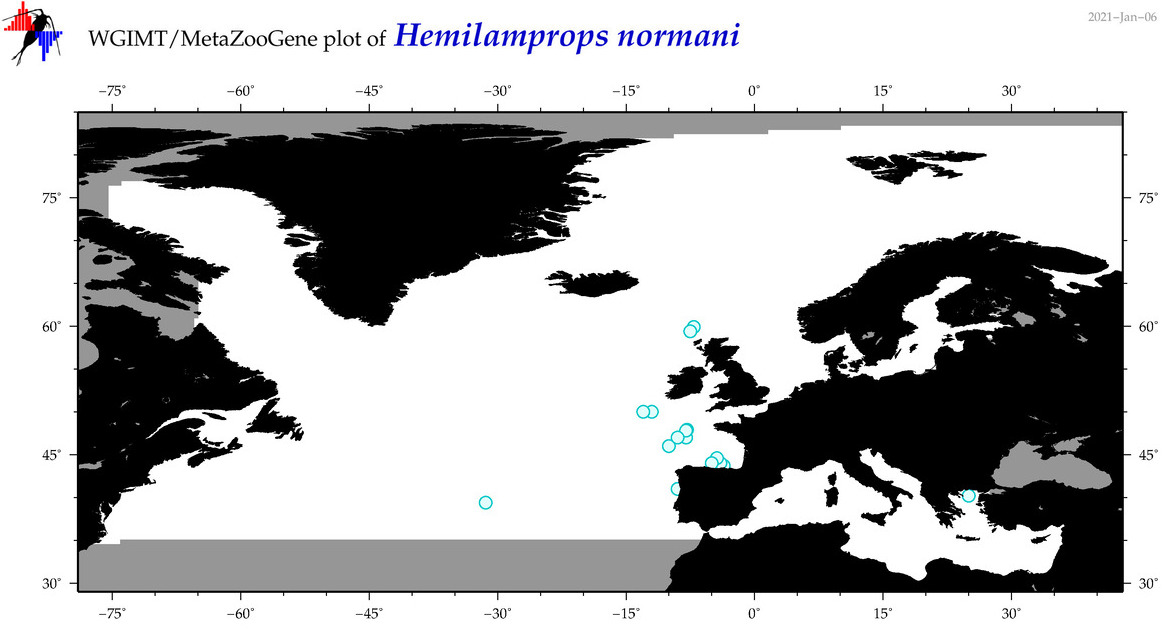
| Hemilamprops normani |
Species
(68) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4053849
R:1:0:0:0 |
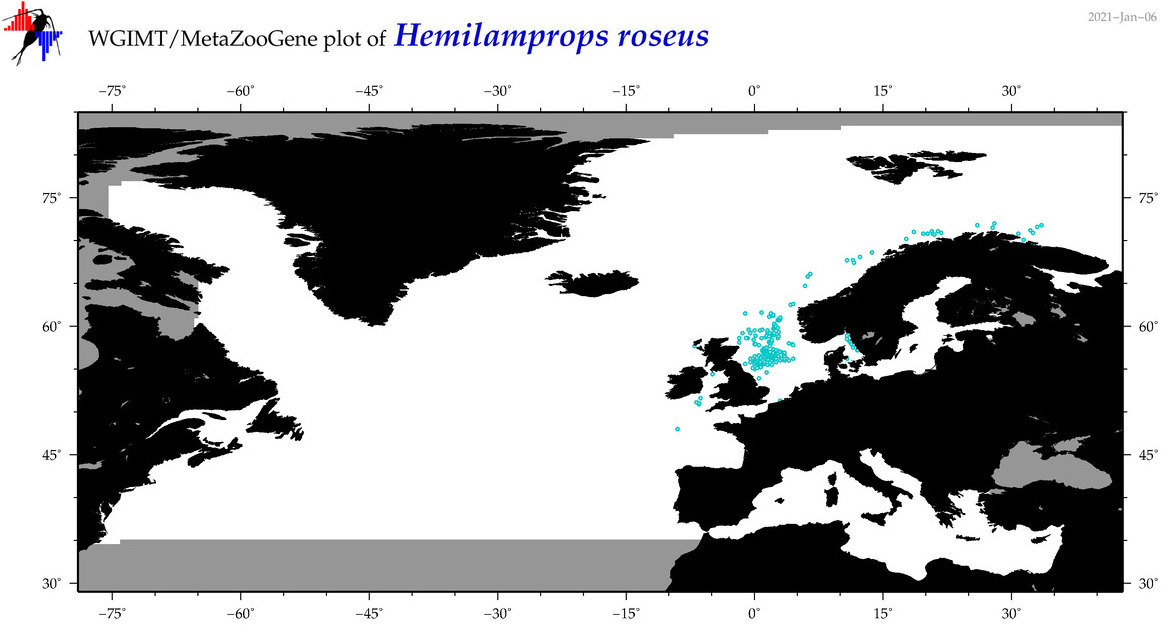
| Hemilamprops roseus |
Species
(69) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4053852
R:1:0:0:0 |
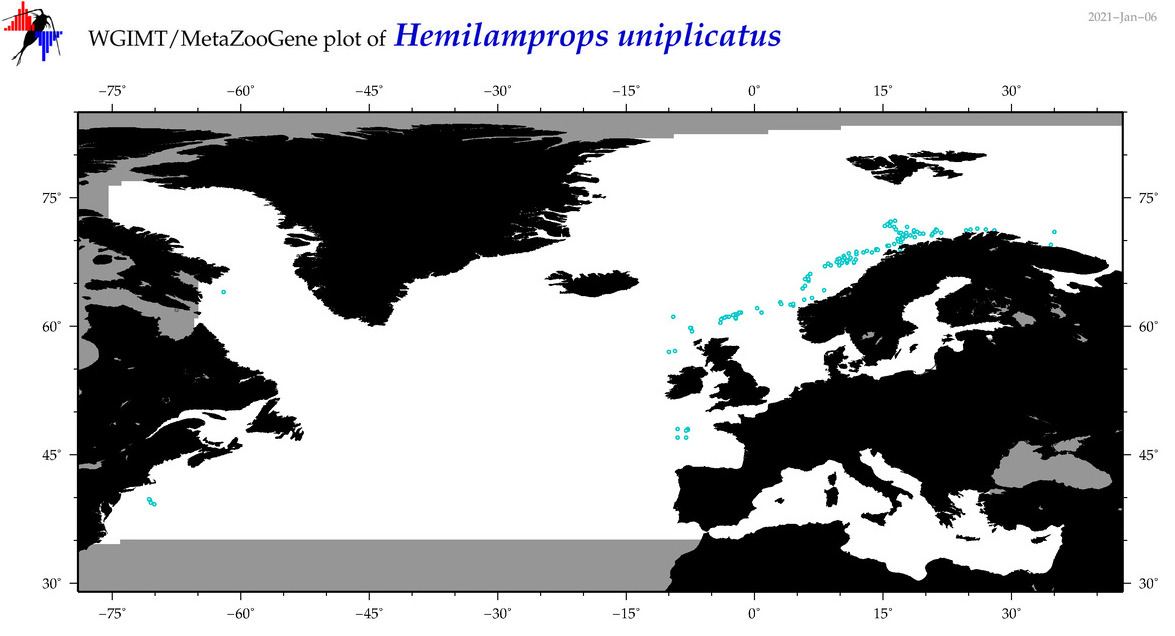
| Hemilamprops uniplicatus |
Species
(70) |
COI
12S
16S
18S
28S
|
COI = 7
12S = 0
16S = 1
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4053854
R:1:0:0:0 |
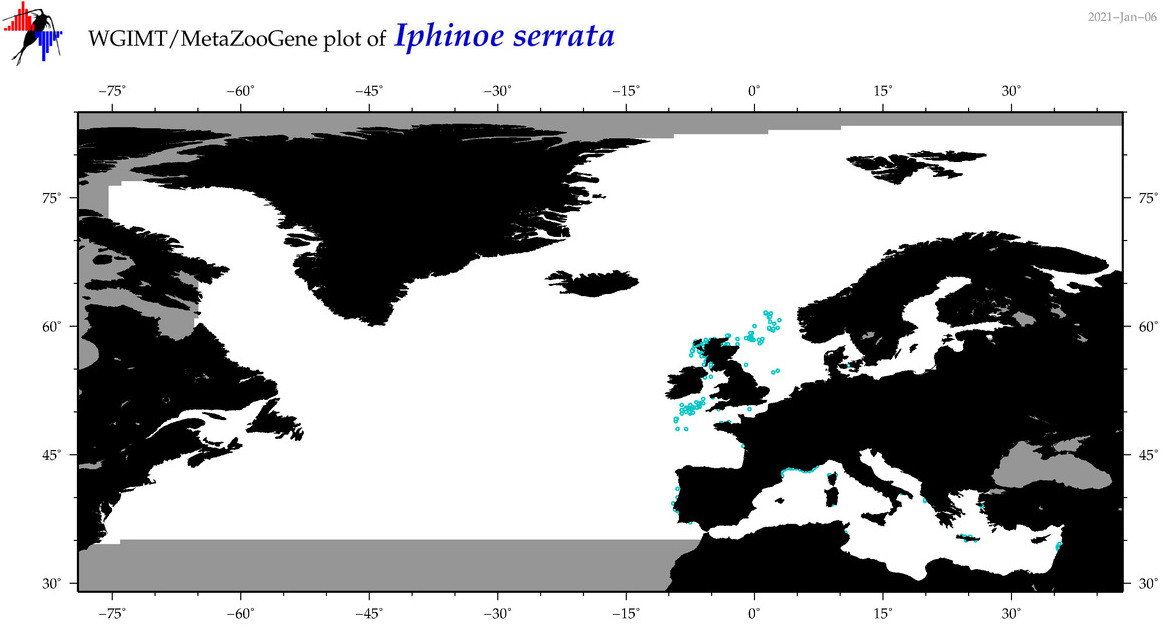
| Iphinoe serrata |
Species
(71) |
COI
12S
16S
18S
28S
|
COI = 2
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4032019
R:1:0:0:0 |
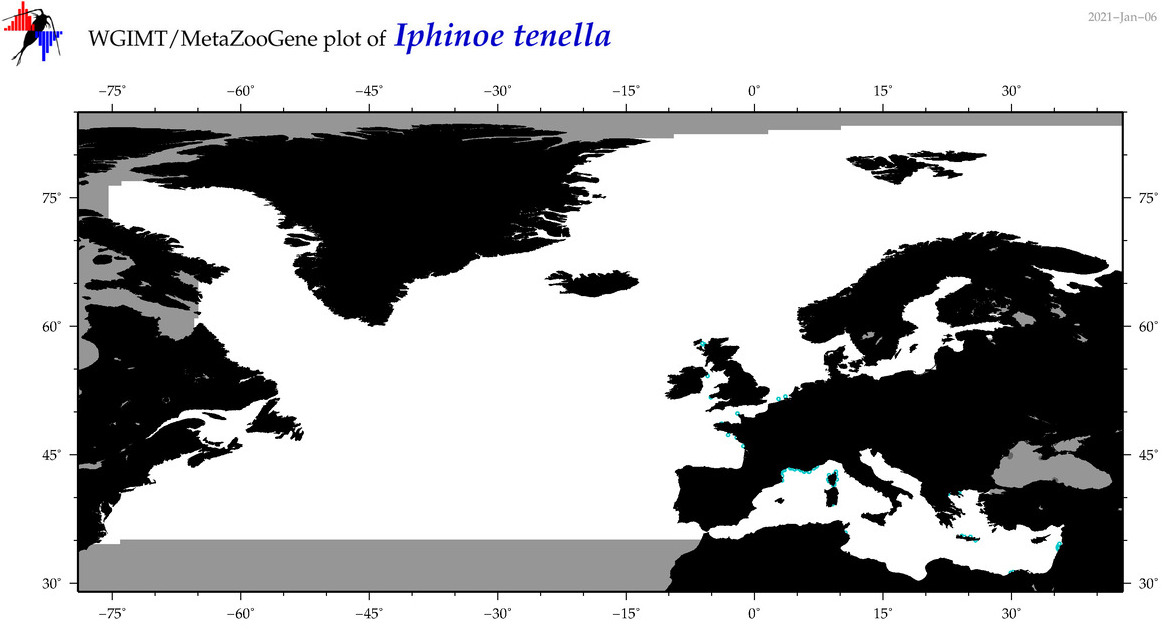
| Iphinoe tenella |
Species
(72) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4054953
R:1:0:0:0 |
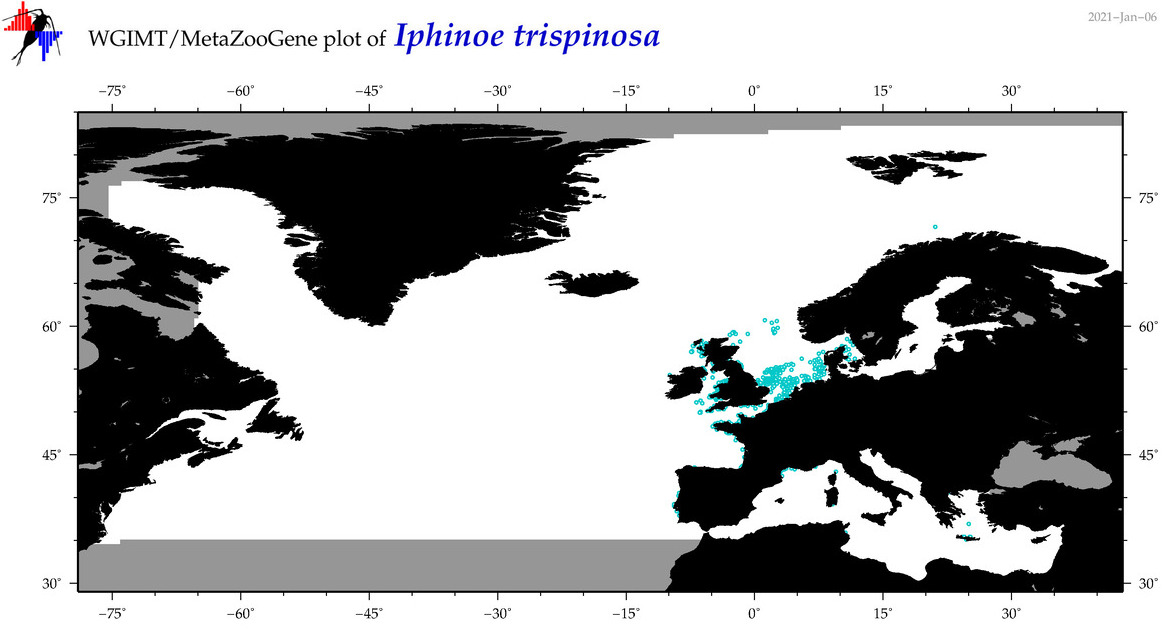
| Iphinoe trispinosa |
Species
(73) |
COI
12S
16S
18S
28S
|
COI = 3
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4034878
R:1:0:0:0 |
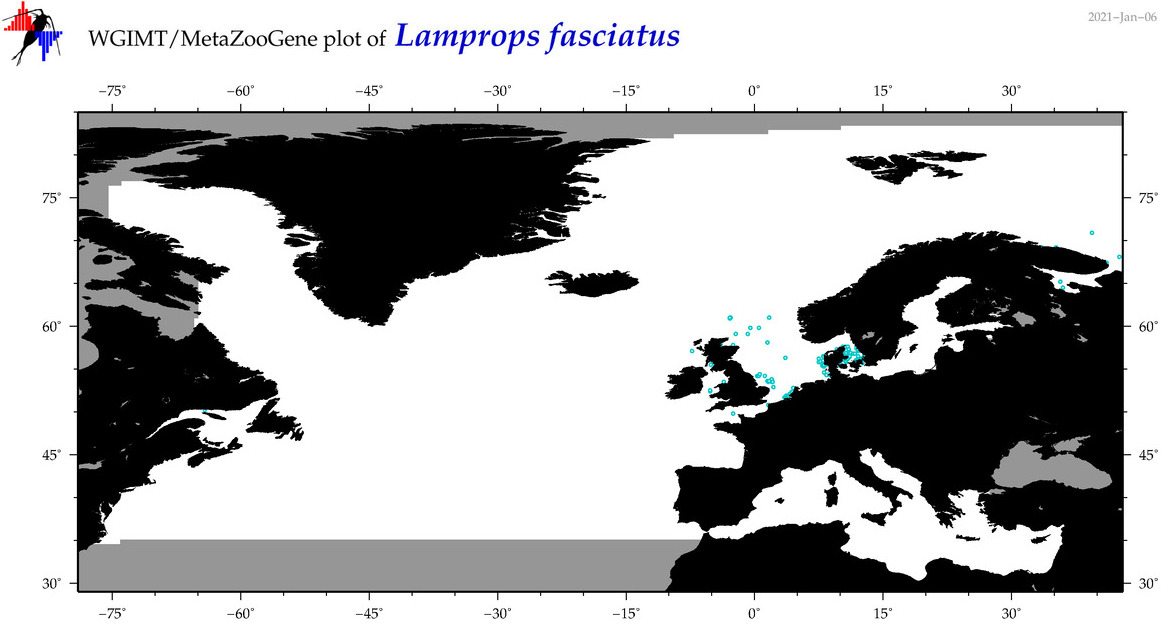
| Lamprops fasciatus |
Species
(74) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4055584
R:1:0:0:0 |
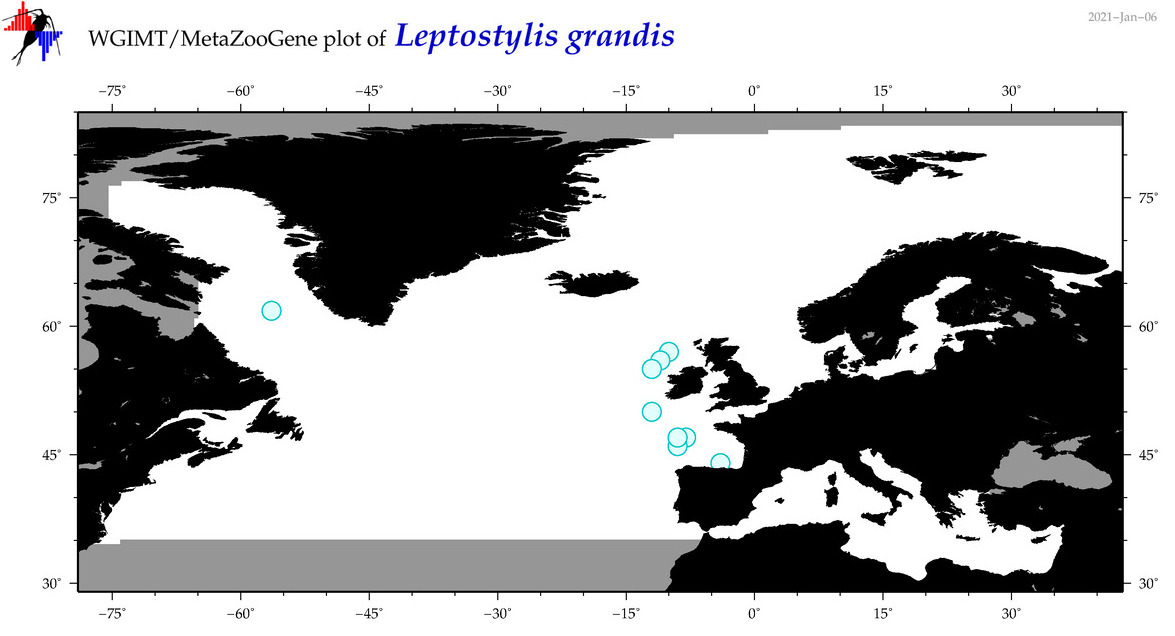
| Leptostylis grandis |
Species
(75) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4056241
R:1:1:0:0 |
| Leptostylis longicaudata |
Species
(76) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4056244
R:1:1:0:0 |
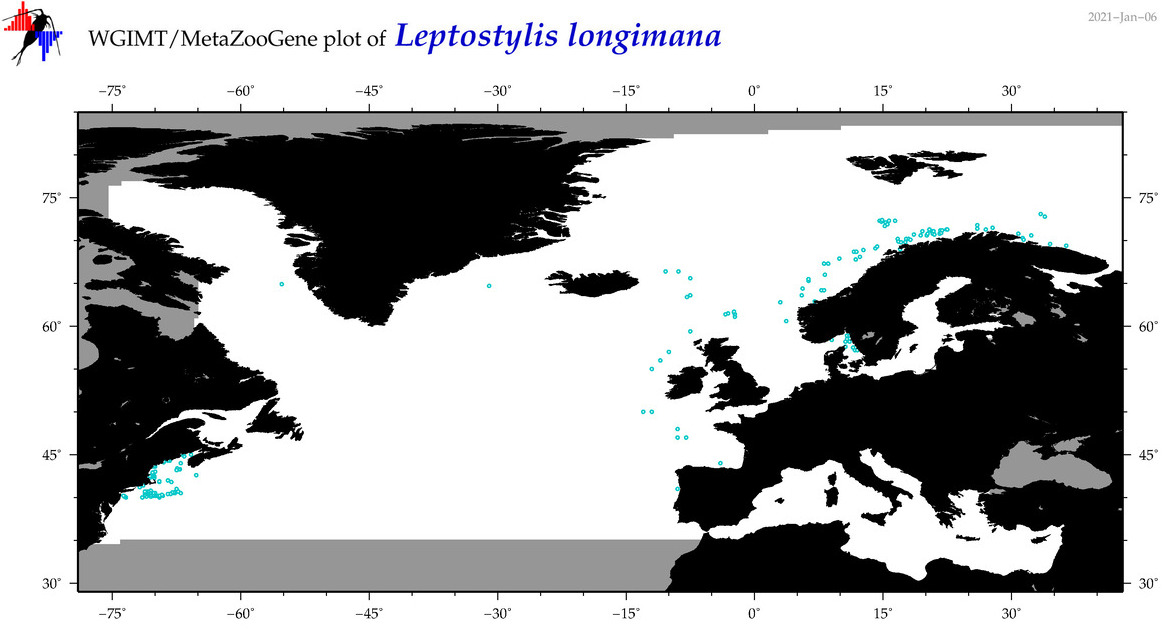
| Leptostylis longimana |
Species
(77) |
COI
12S
16S
18S
28S
|
COI = 2
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4056245
R:1:1:0:0 |
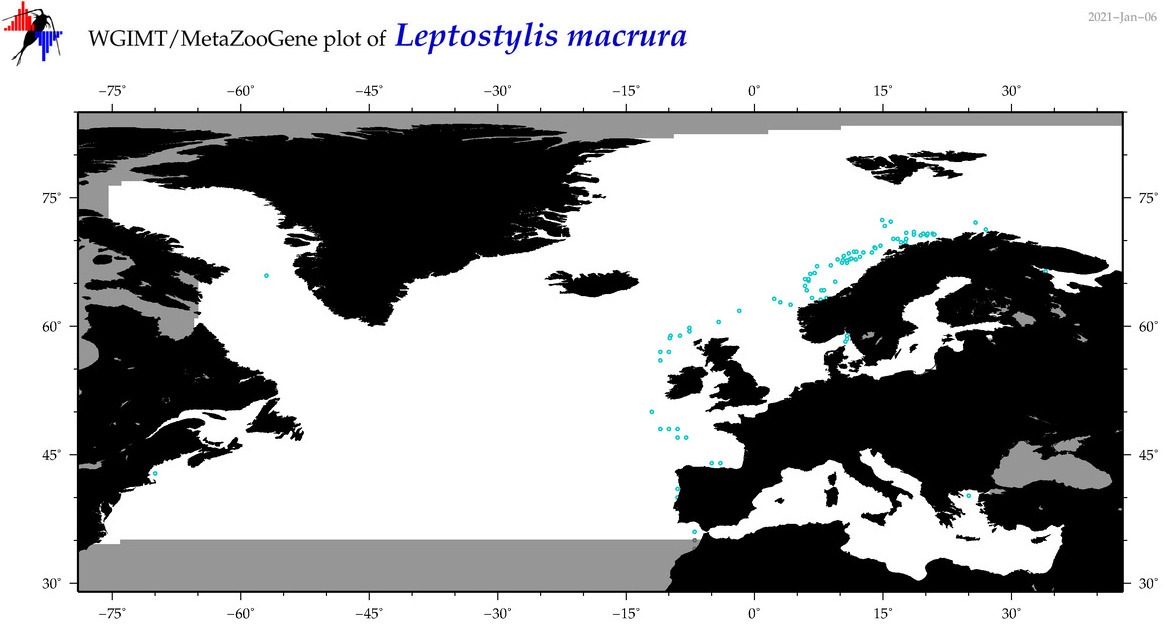
| Leptostylis macrura |
Species
(78) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4056246
R:1:1:0:0 |
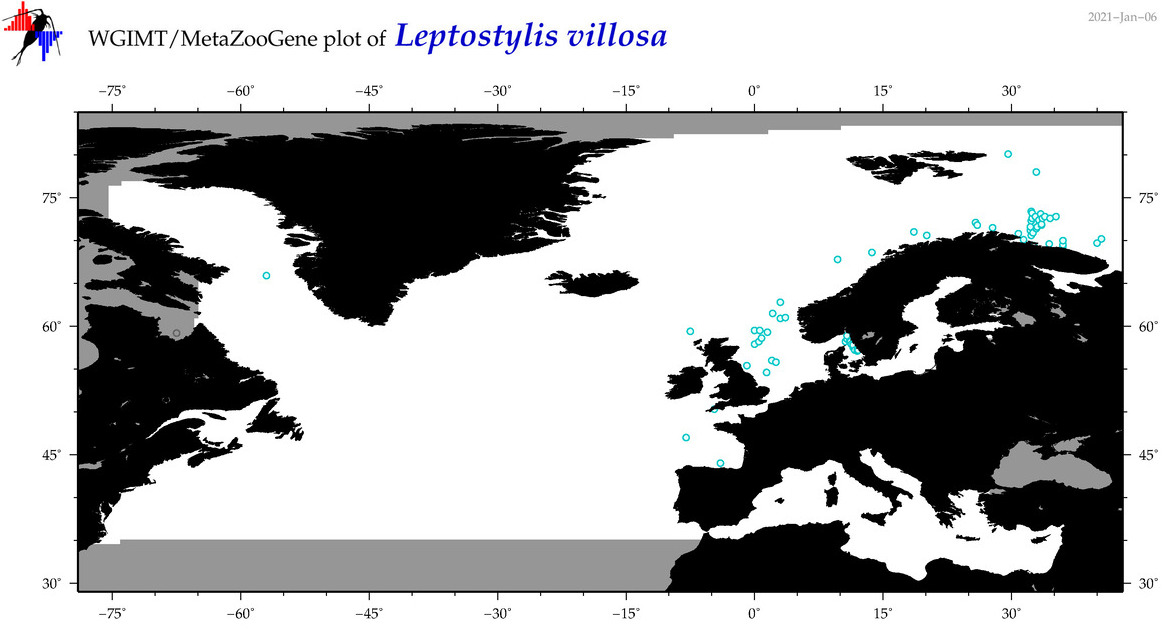
| Leptostylis villosa |
Species
(79) |
COI
12S
16S
18S
28S
|
COI = 7
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4056253
R:1:1:0:0 |
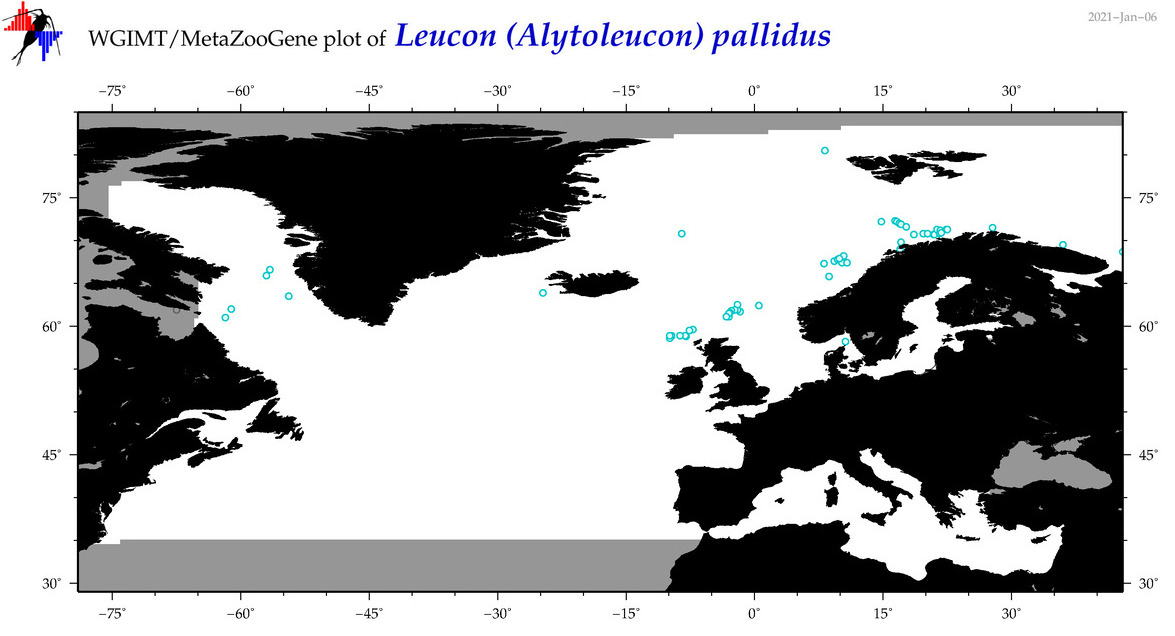
| Leucon (Alytoleucon) pallidus |
Species
(80) |
COI
12S
16S
18S
28S
|
COI = 2
12S = 0
16S = 7
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091486
R:1:1:0:0 |
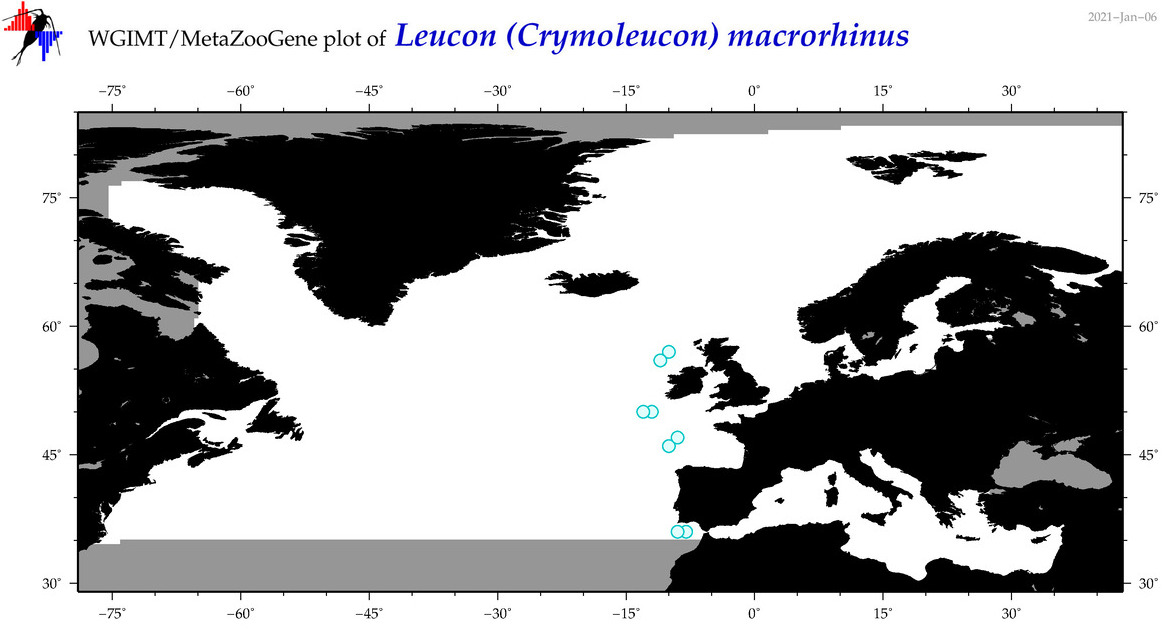
| Leucon (Crymoleucon) macrorhinus |
Species
(81) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091487
R:1:1:0:0 |
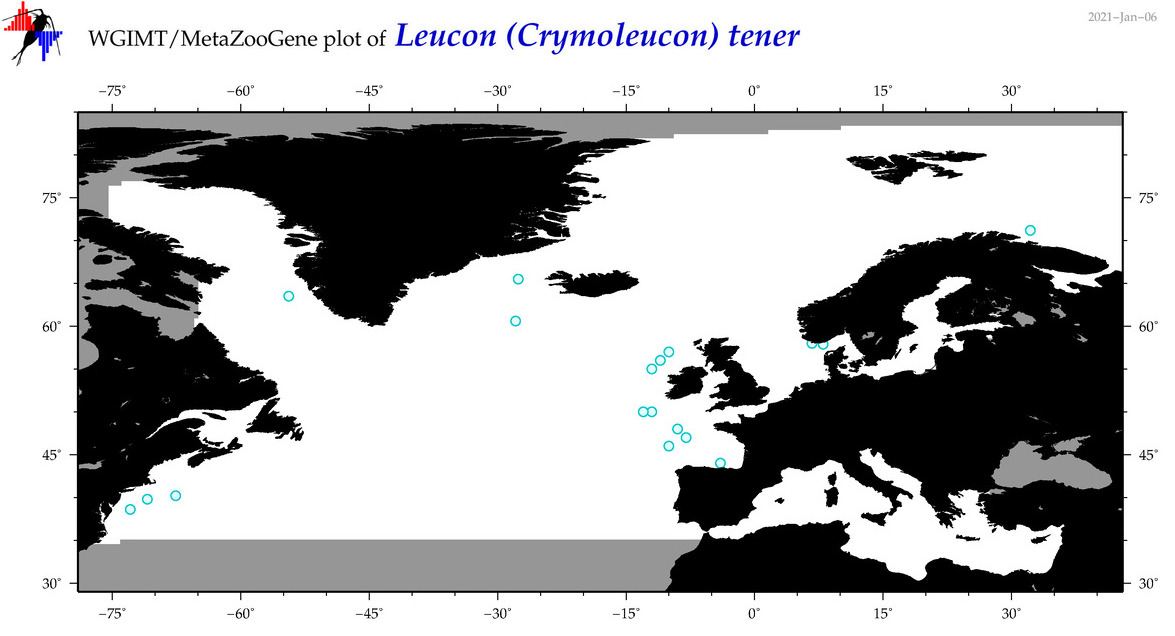
| Leucon (Crymoleucon) tener |
Species
(82) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091488
R:1:1:0:0 |
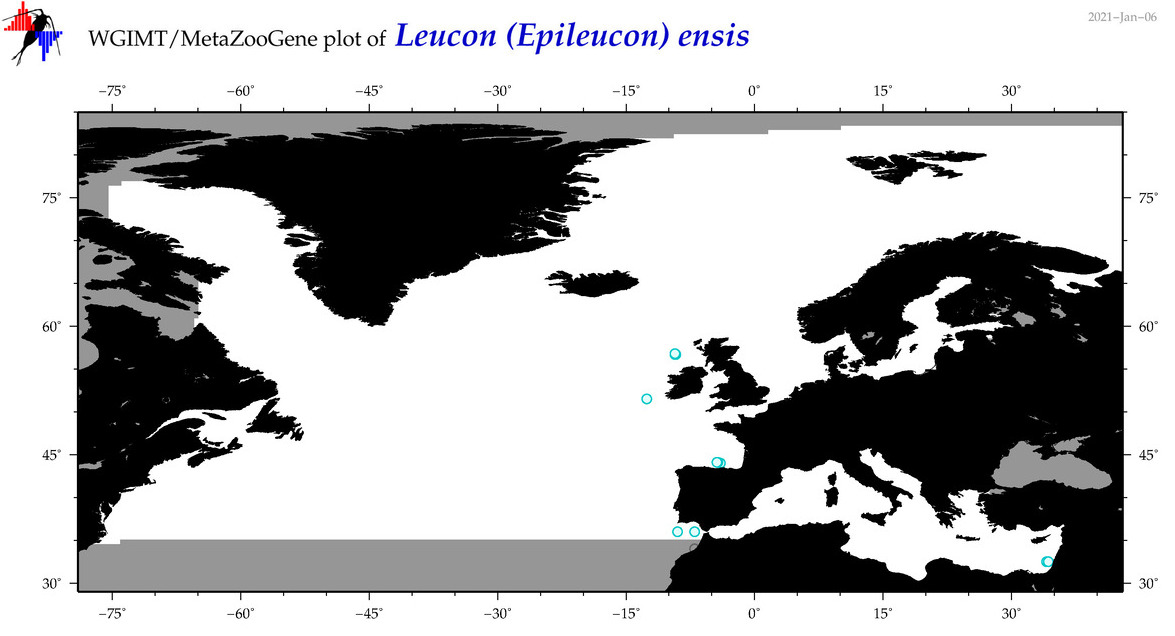
| Leucon (Epileucon) ensis |
Species
(83) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094851
R:1:1:0:0 |
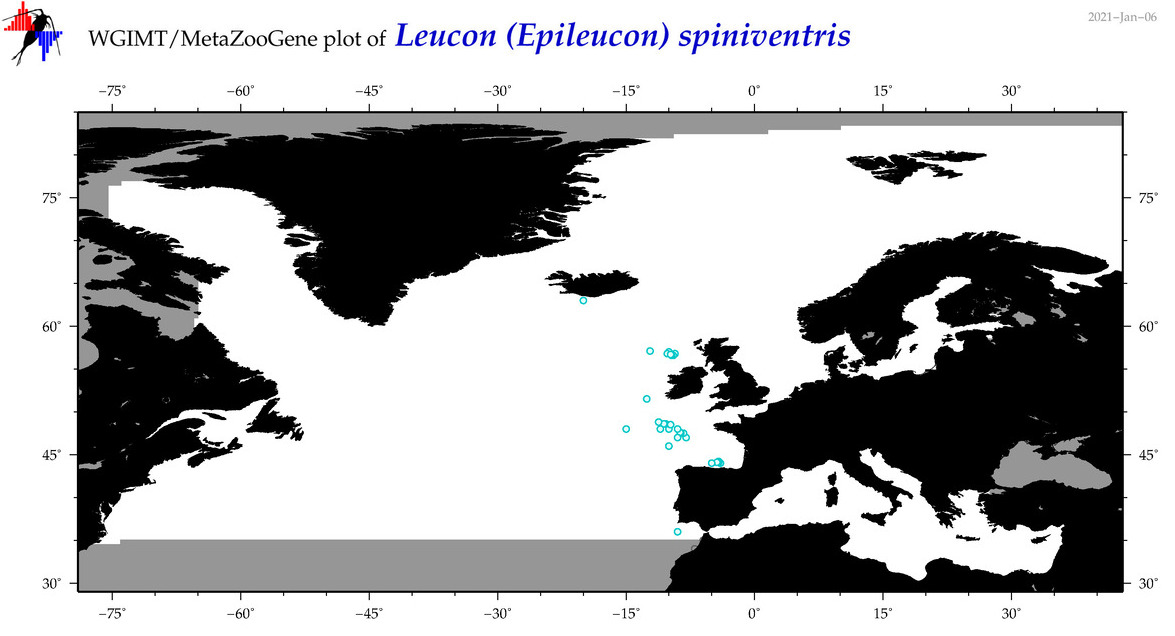
| Leucon (Epileucon) spiniventris |
Species
(84) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094855
R:1:1:0:0 |
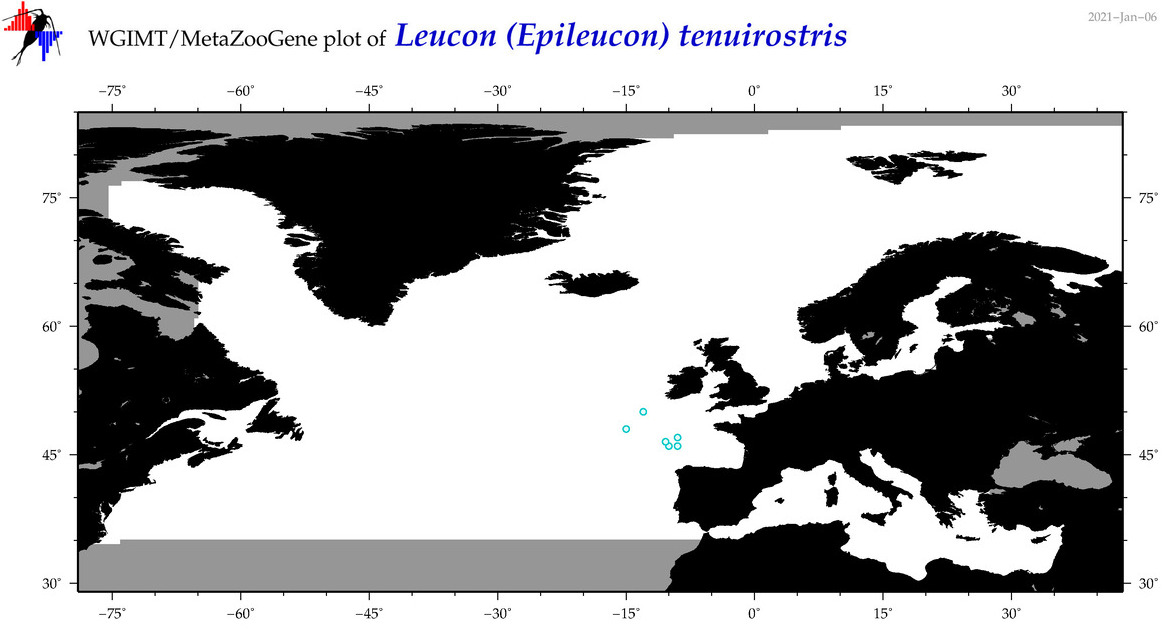
| Leucon (Epileucon) tenuirostris |
Species
(85) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091490
R:1:1:0:0 |
| Leucon hanseni |
Species
(86) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4056287
R:1:1:0:0 |
| Leucon holti |
Species
(87) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4056288
R:1:1:0:0 |
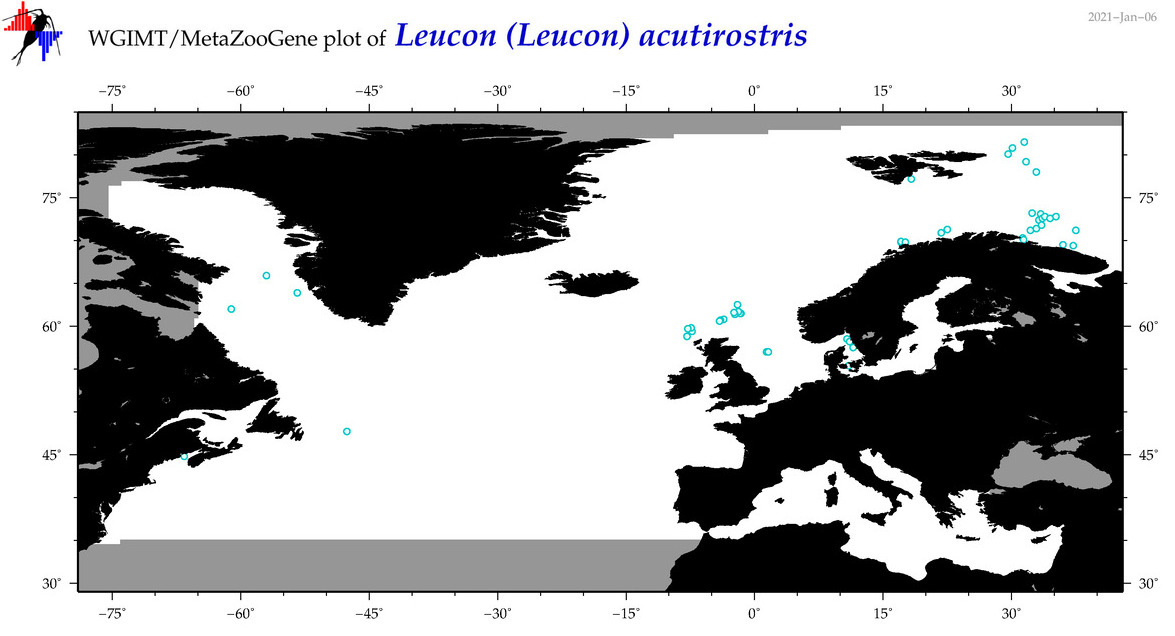
| Leucon (Leucon) acutirostris |
Species
(88) |
COI
12S
16S
18S
28S
|
COI = 2
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091491
R:1:1:0:0 |
| Leucon (Leucon) fulvus |
Species
(89) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091492
R:1:1:0:0 |
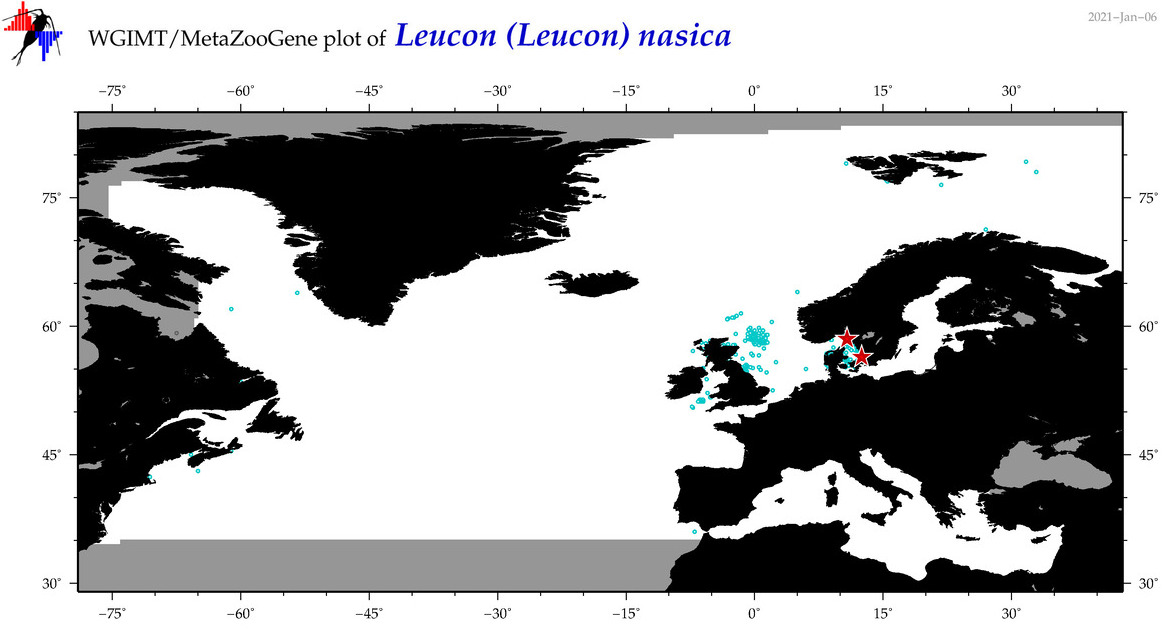
| Leucon (Leucon) nasica |
Species
(90) |
COI
12S
16S
18S
28S
|
COI = 8
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091494
R:1:1:0:0 |
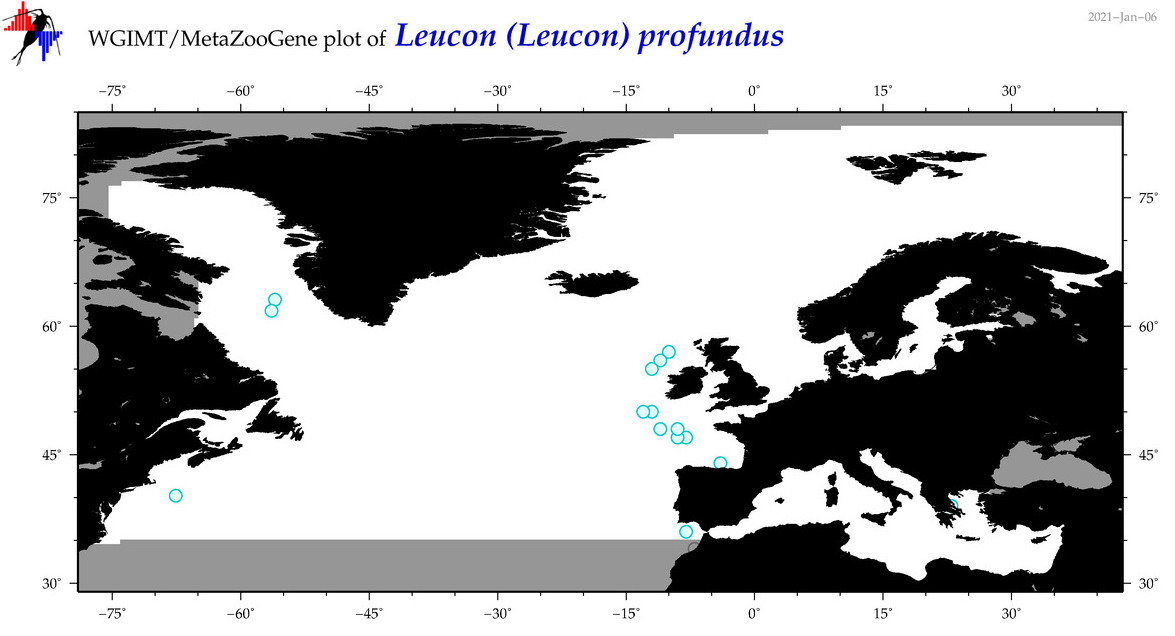
| Leucon (Leucon) profundus |
Species
(91) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094866
R:1:1:0:0 |
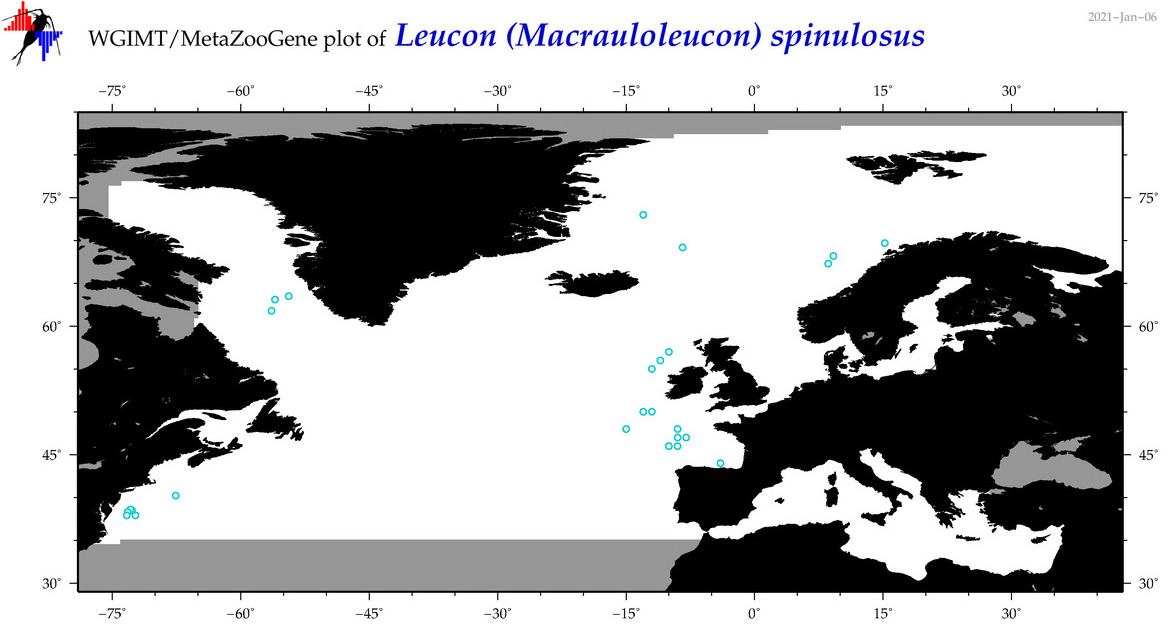
| Leucon (Macrauloleucon) spinulosus |
Species
(92) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091500
R:1:1:0:0 |
| Leucon sarsi |
Species
(93) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4056292
R:1:1:0:0 |
| Makrokylindrus (Adiastylis) alleni |
Species
(94) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094805
R:1:1:0:0 |
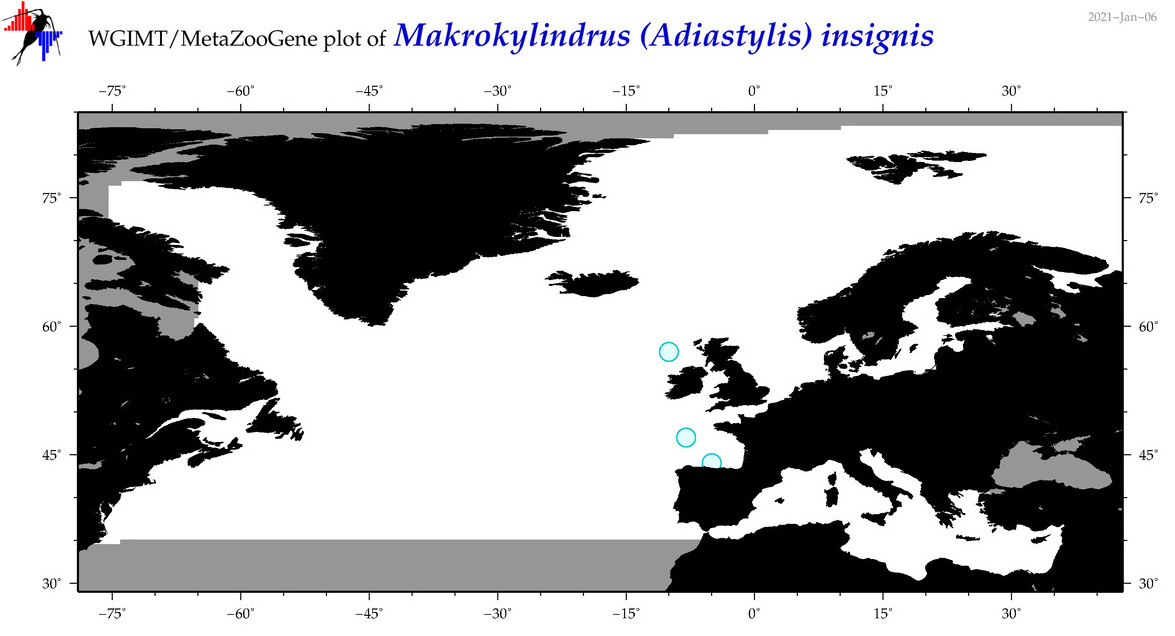
| Makrokylindrus (Adiastylis) insignis |
Species
(95) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091478
R:1:1:0:0 |
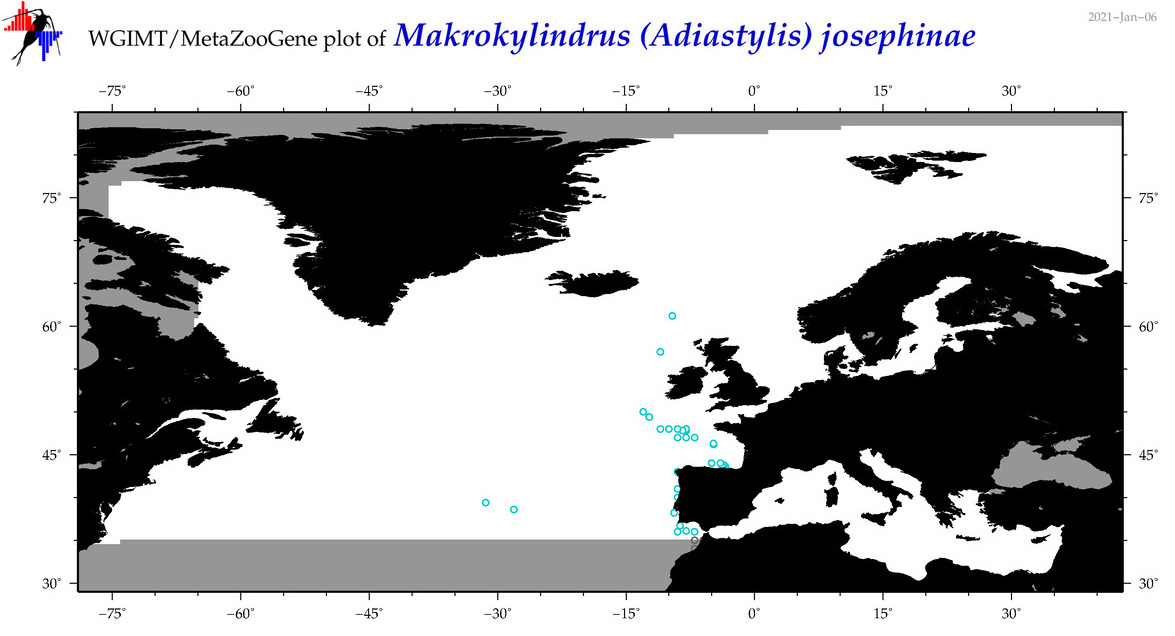
| Makrokylindrus (Adiastylis) josephinae |
Species
(96) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091479
R:1:1:0:0 |
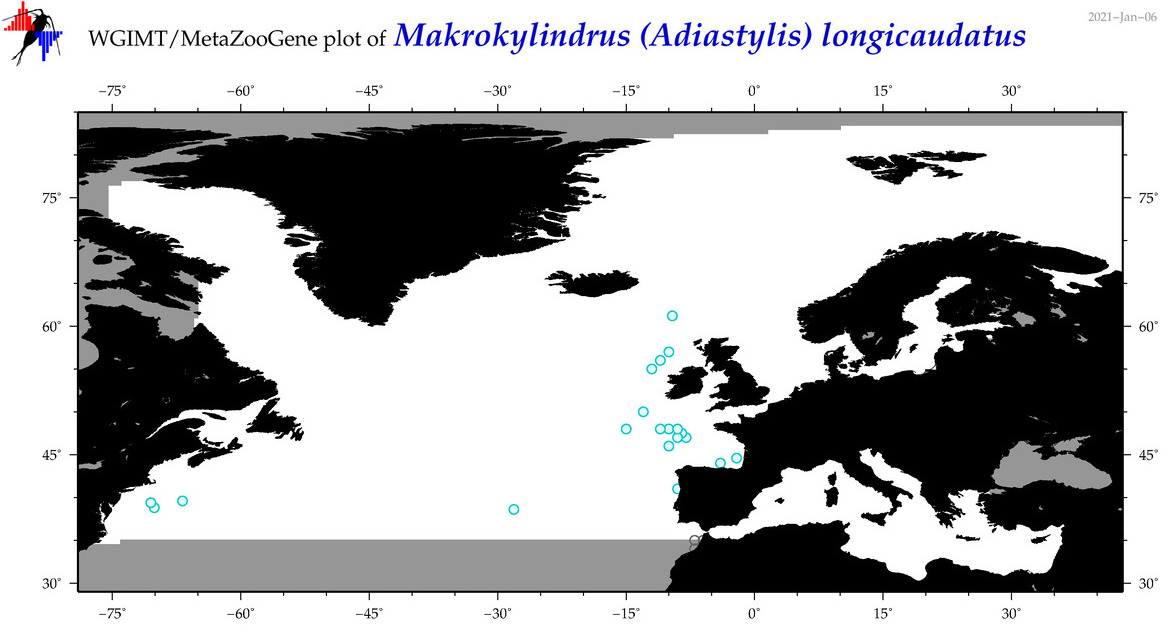
| Makrokylindrus (Adiastylis) longicaudatus |
Species
(97) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091480
R:1:1:0:0 |
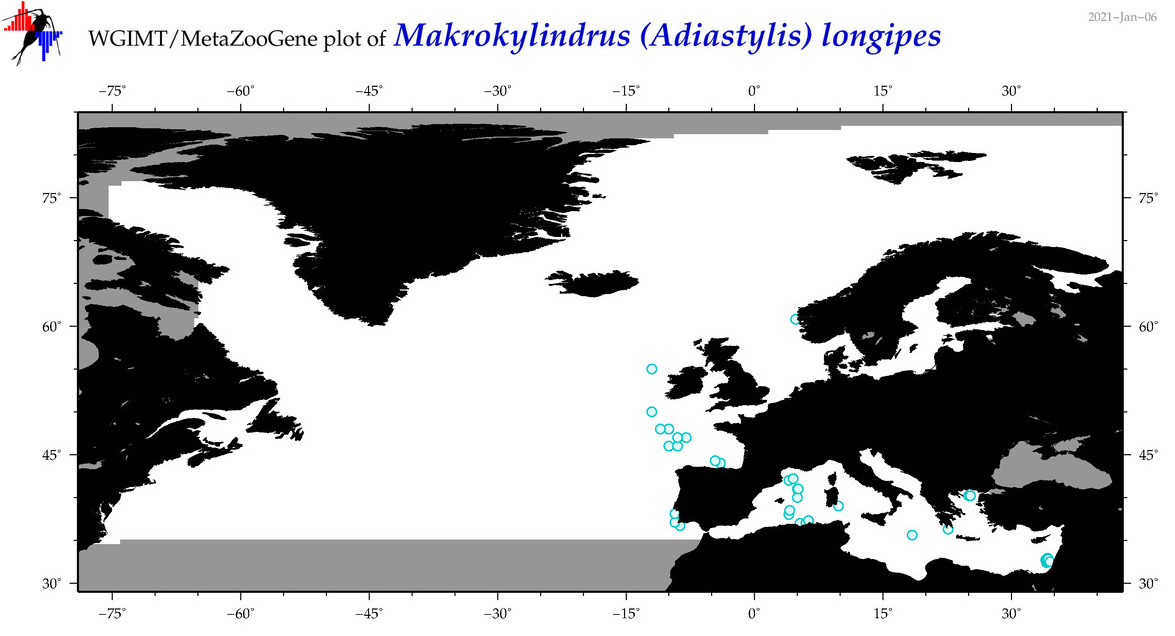
| Makrokylindrus (Adiastylis) longipes |
Species
(98) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091481
R:1:1:0:0 |
| Makrokylindrus (Adiastylis) myriamae |
Species
(99) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
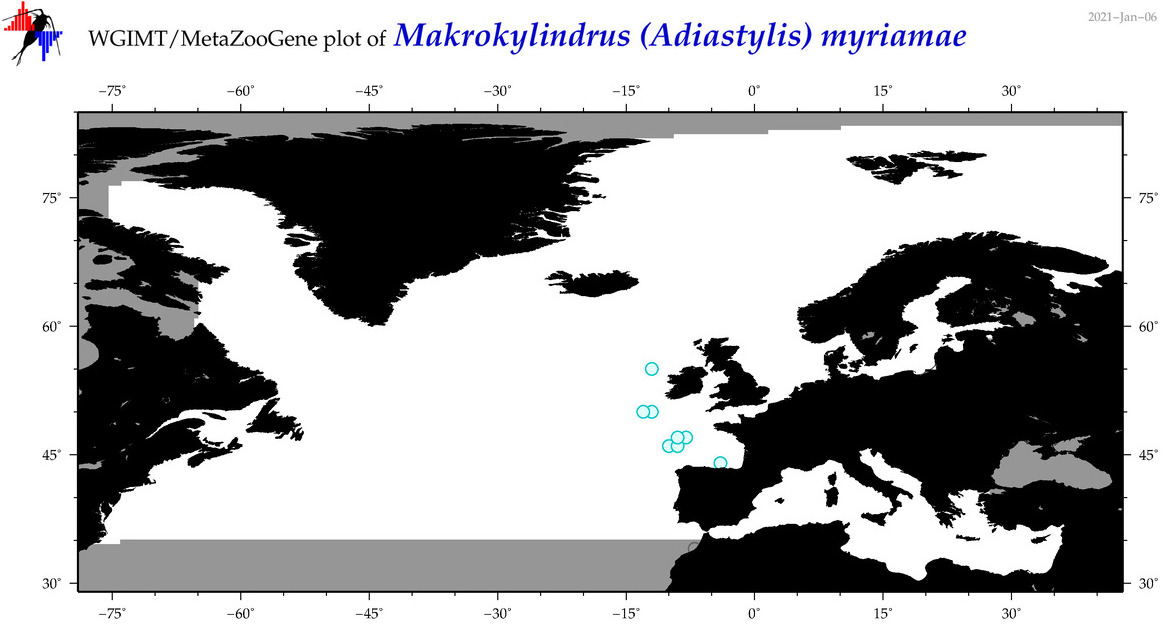 |
accepted
T4094822
R:1:1:0:0 |
| Makrokylindrus (Adiastylis) tubulicauda |
Species
(100) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
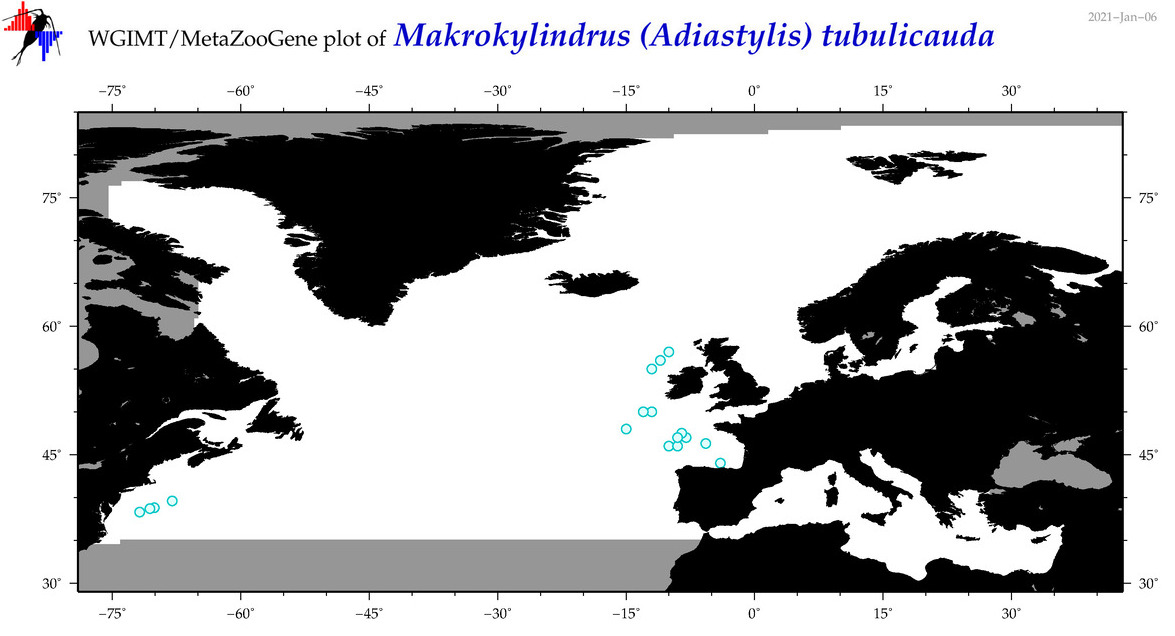 |
accepted
T4091482
R:1:1:0:0 |
| Makrokylindrus (Makrokylindrus) dubius |
Species
(101) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
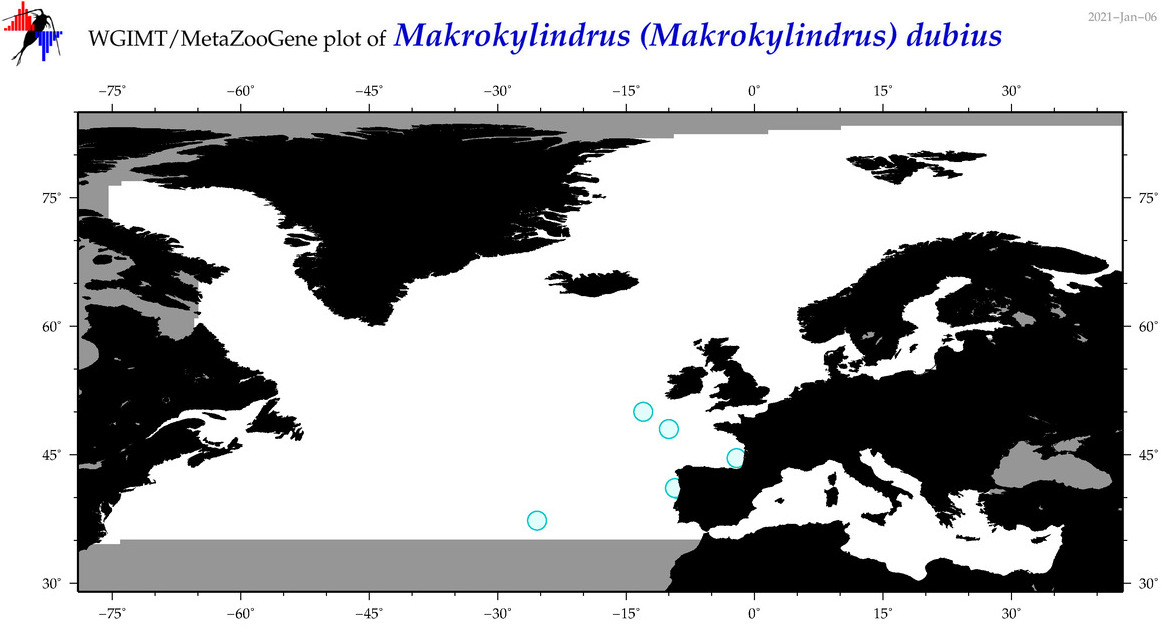 |
accepted
T4091484
R:1:1:0:0 |
| Makrokylindrus (Makrokylindrus) mystacinus |
Species
(102) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4094836
R:1:1:0:0 |
| Makrokylindrus (Makrokylindrus) spiniventris |
Species
(103) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
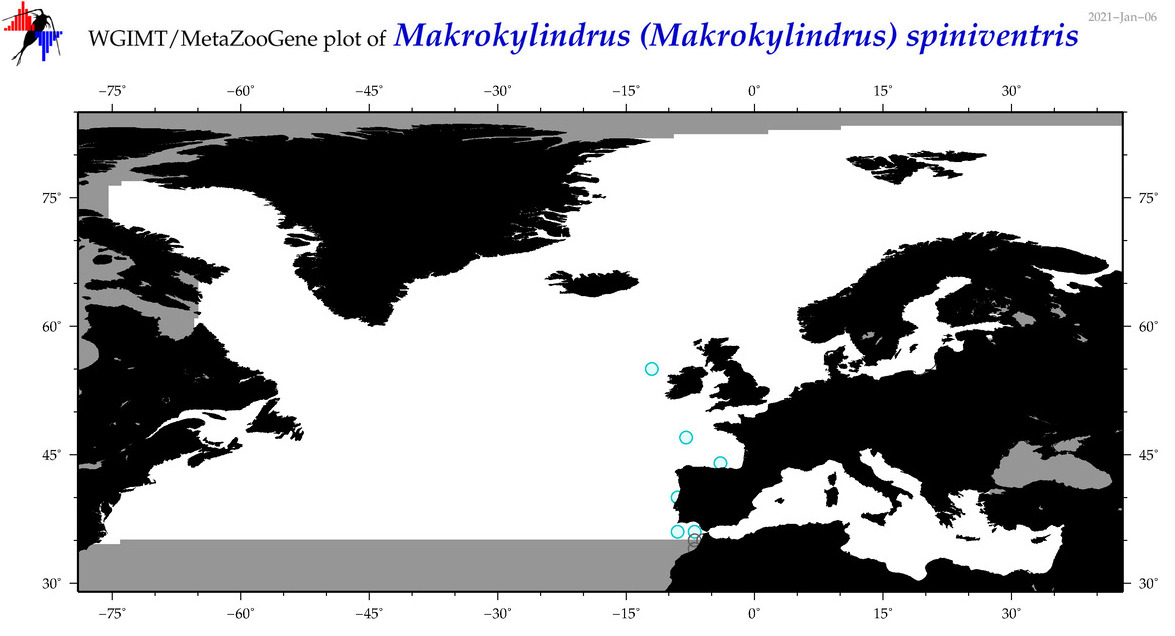 |
accepted
T4091485
R:1:1:0:0 |
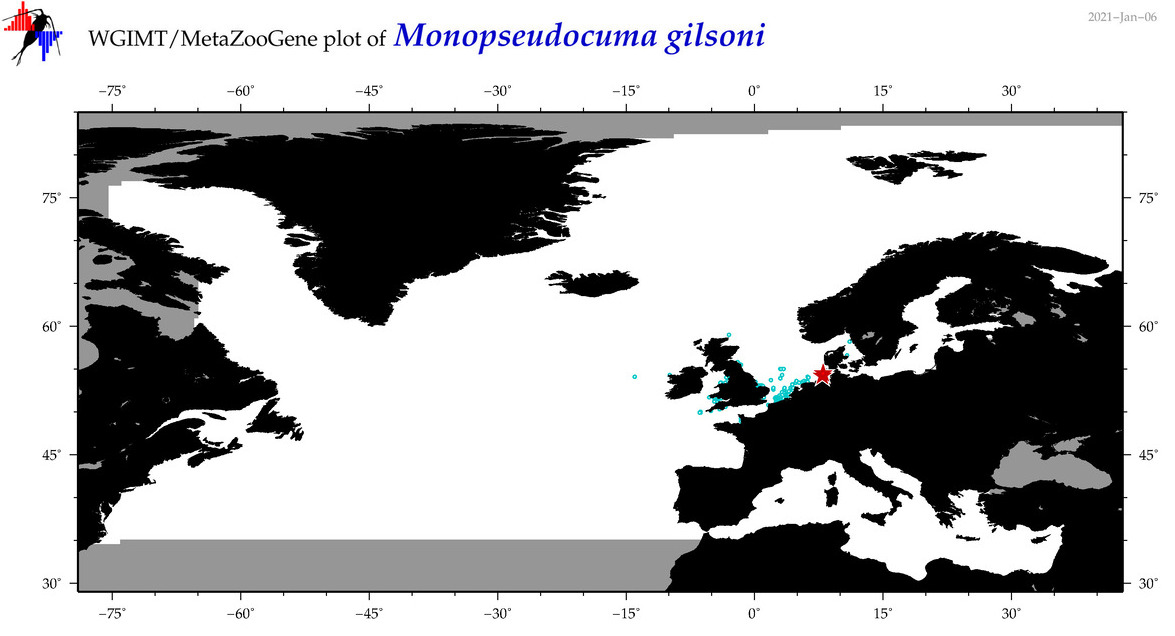
| Monopseudocuma gilsoni |
Species
(104) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 1
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4013106
R:1:0:0:0 |
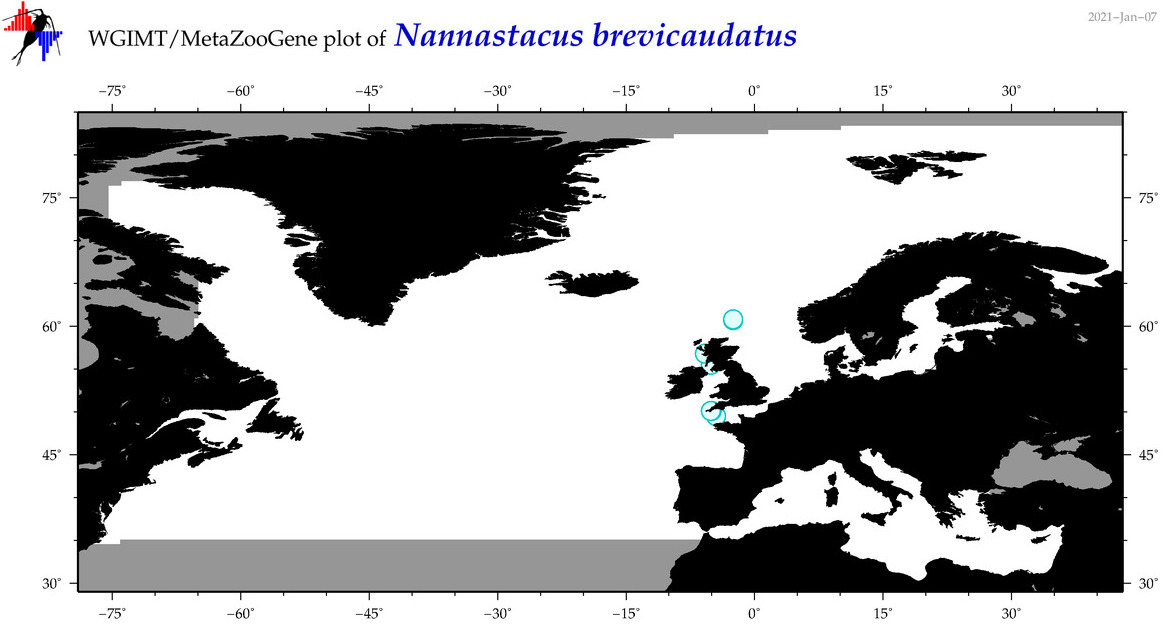
| Nannastacus brevicaudatus |
Species
(105) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4059690
R:1:0:0:0 |
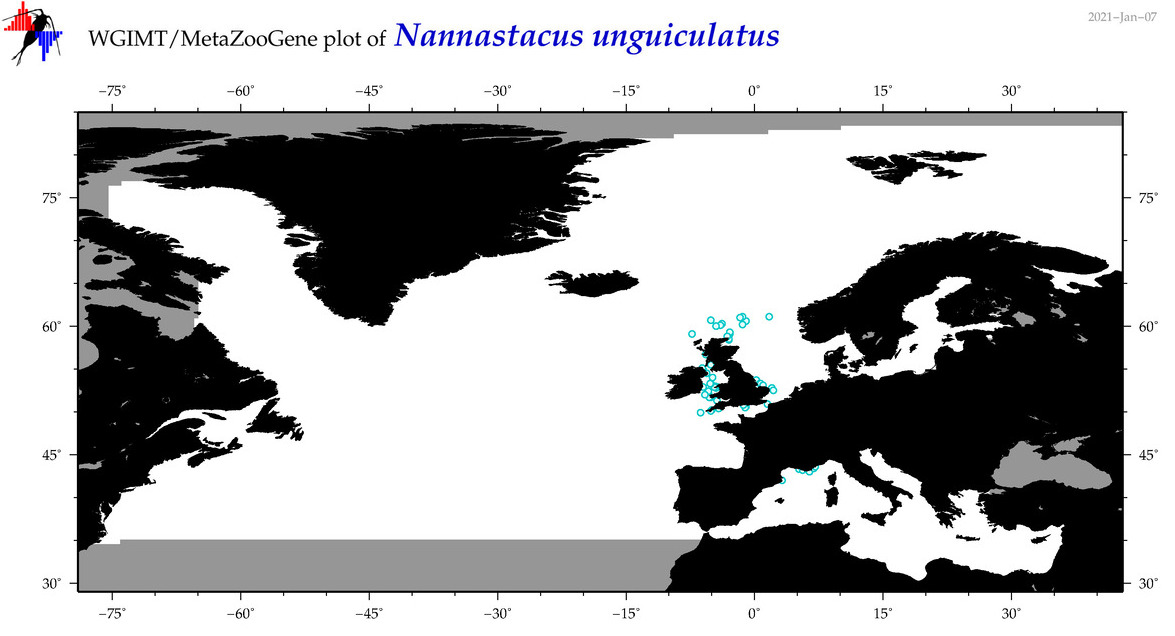
| Nannastacus unguiculatus |
Species
(106) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4059700
R:1:0:0:0 |
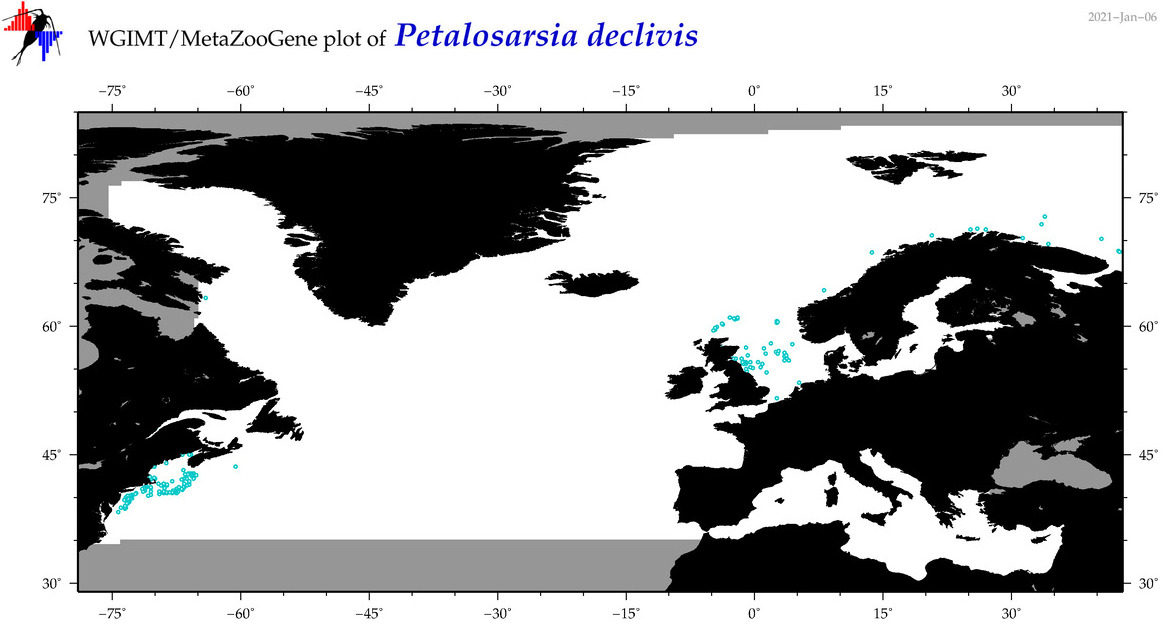
| Petalosarsia declivis |
Species
(107) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4063491
R:1:0:0:0 |
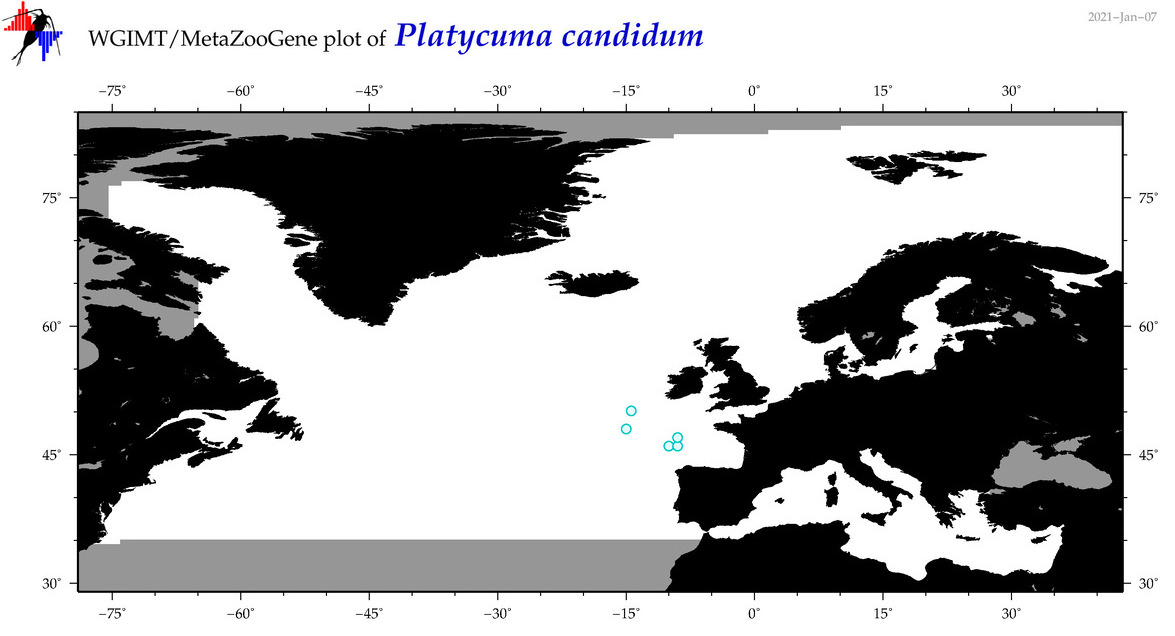
| Platycuma candidum |
Species
(108) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064205
R:1:0:0:0 |
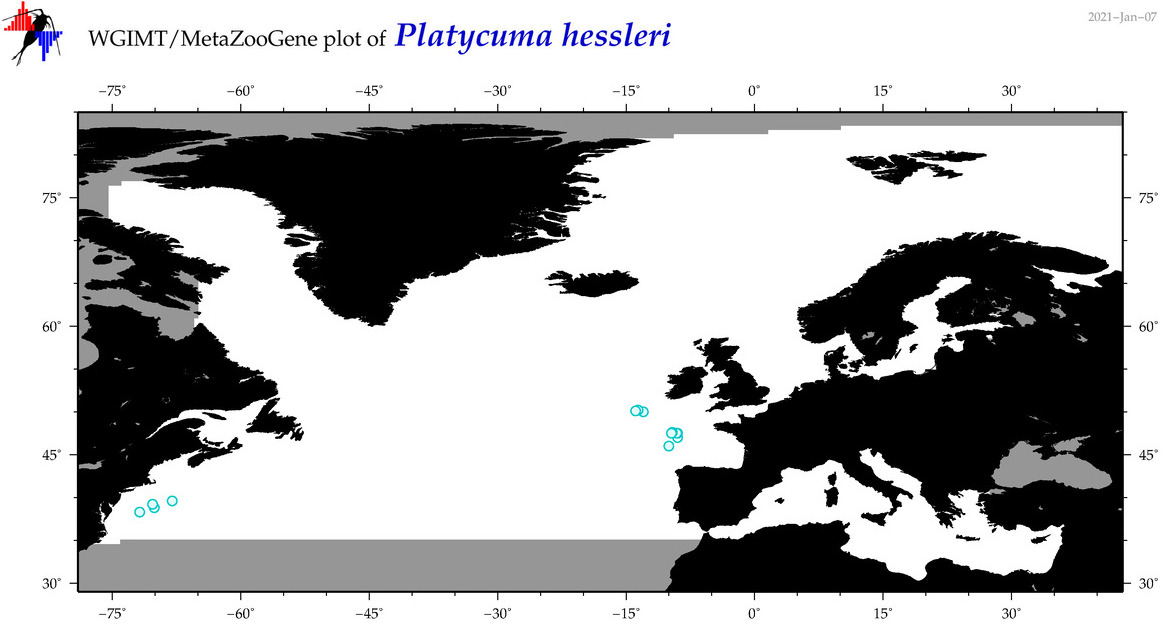
| Platycuma hessleri |
Species
(109) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064206
R:1:0:0:0 |
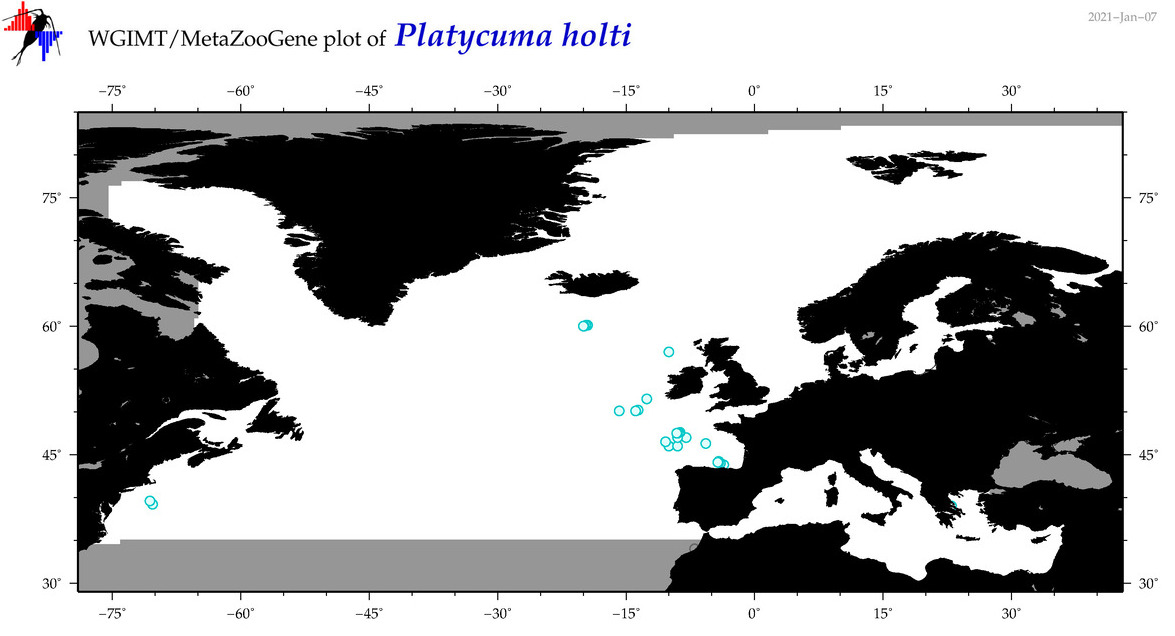
| Platycuma holti |
Species
(110) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064207
R:1:0:0:0 |
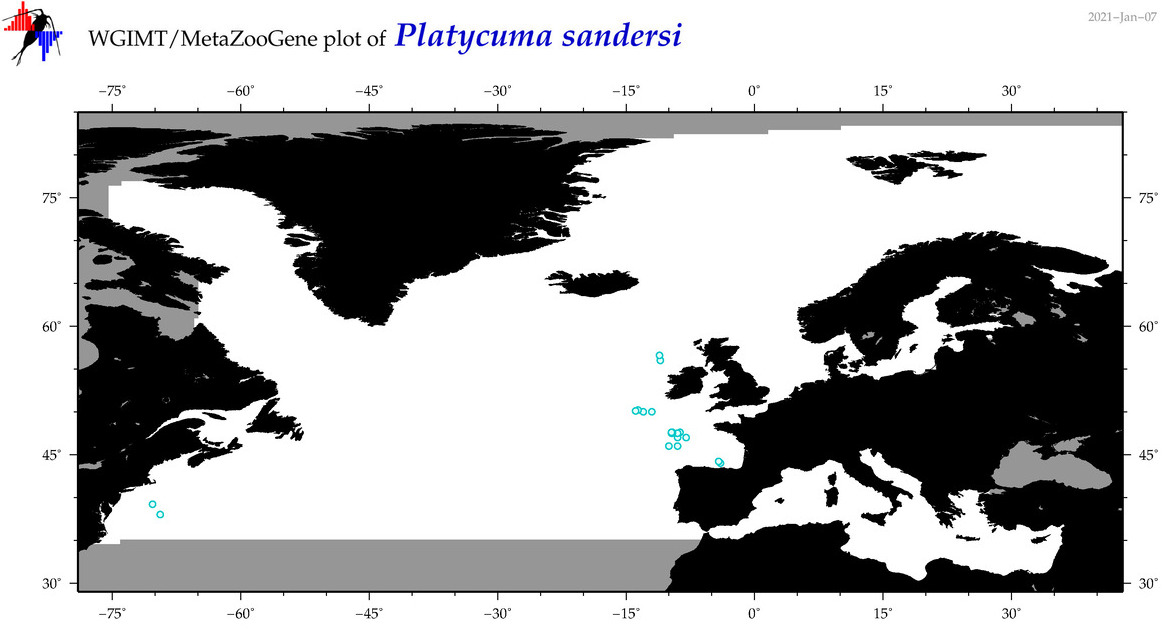
| Platycuma sandersi |
Species
(111) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064210
R:1:0:0:0 |
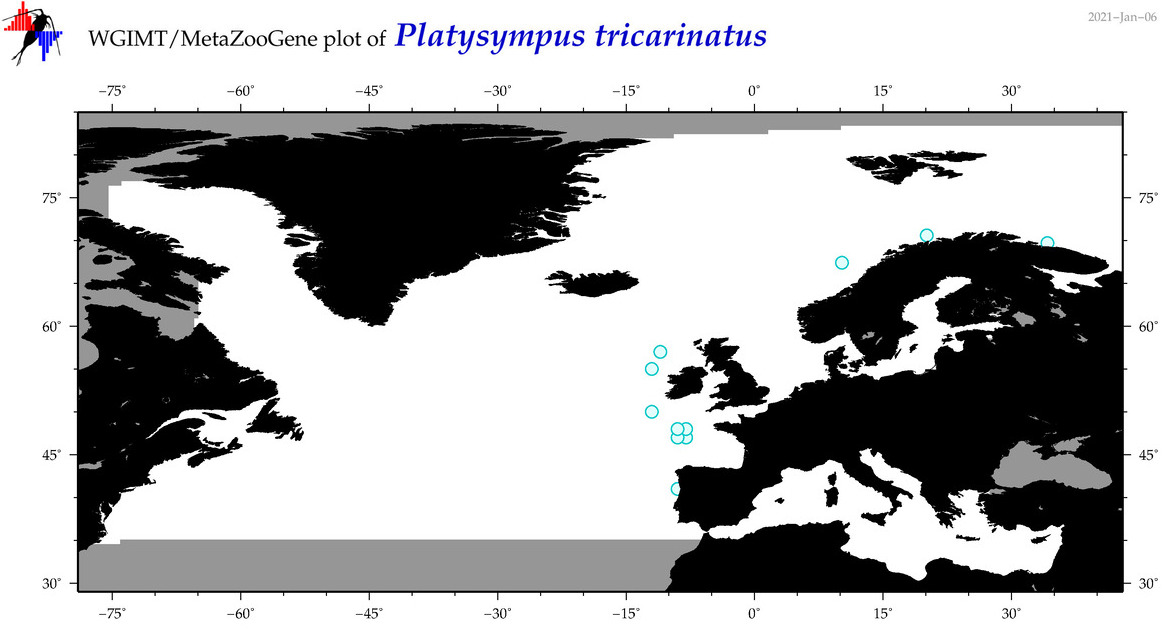
| Platysympus tricarinatus |
Species
(112) |
COI
12S
16S
18S
28S
|
COI = 6
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064263
R:1:0:0:0 |
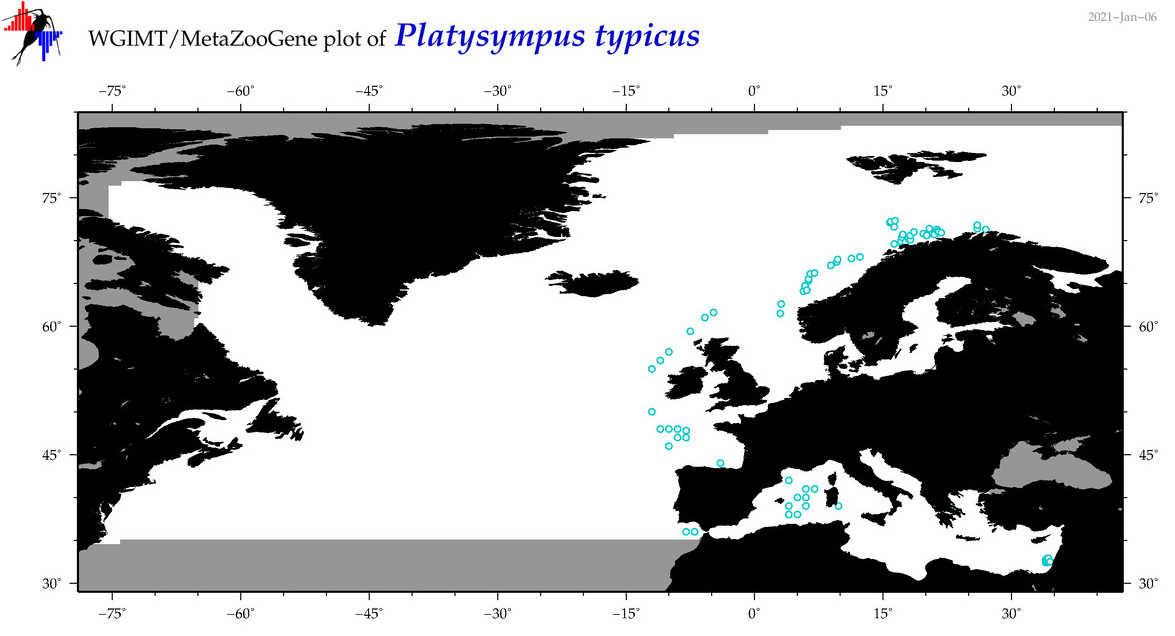
| Platysympus typicus |
Species
(113) |
COI
12S
16S
18S
28S
|
COI = 4
12S = 0
16S = 2
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064264
R:1:0:0:0 |
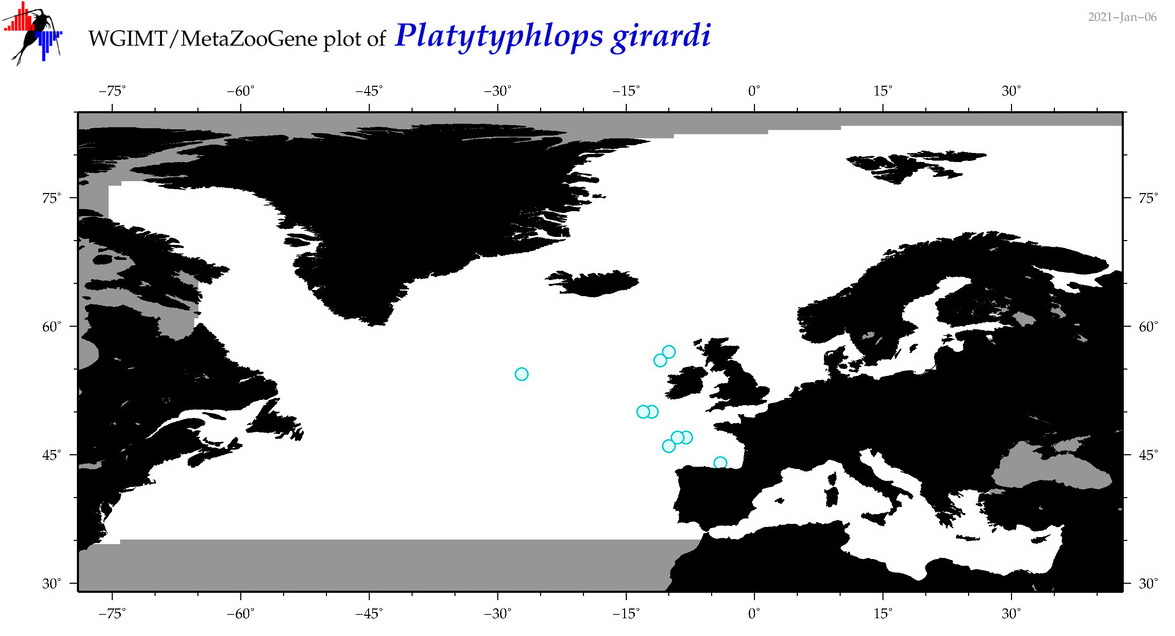
| Platytyphlops girardi |
Species
(114) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064276
R:1:0:0:0 |
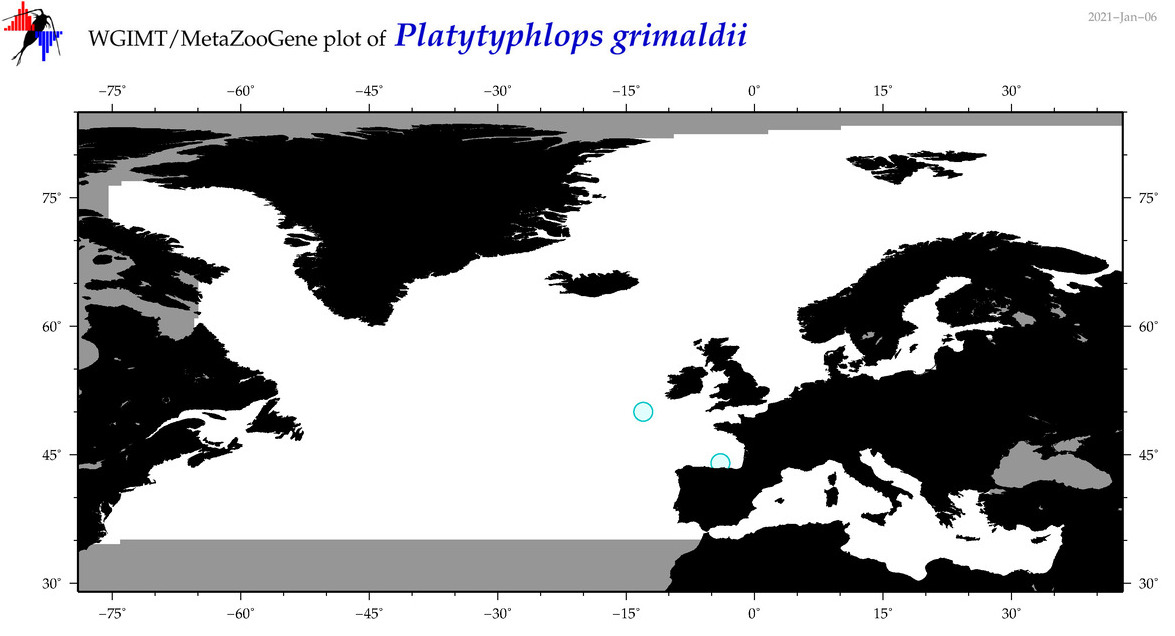
| Platytyphlops grimaldii |
Species
(115) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064277
R:1:0:0:0 |
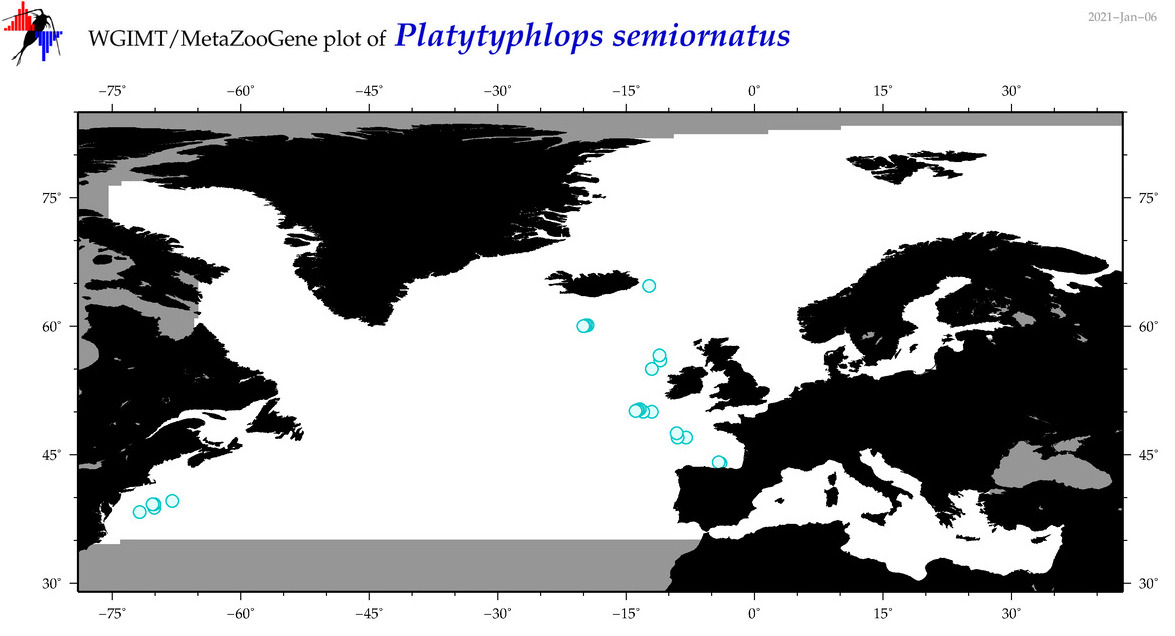
| Platytyphlops semiornatus |
Species
(116) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4064284
R:1:0:0:0 |
| Platytyphlops tuberculatus |
Species
(117) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4064285
R:1:0:0:0 |
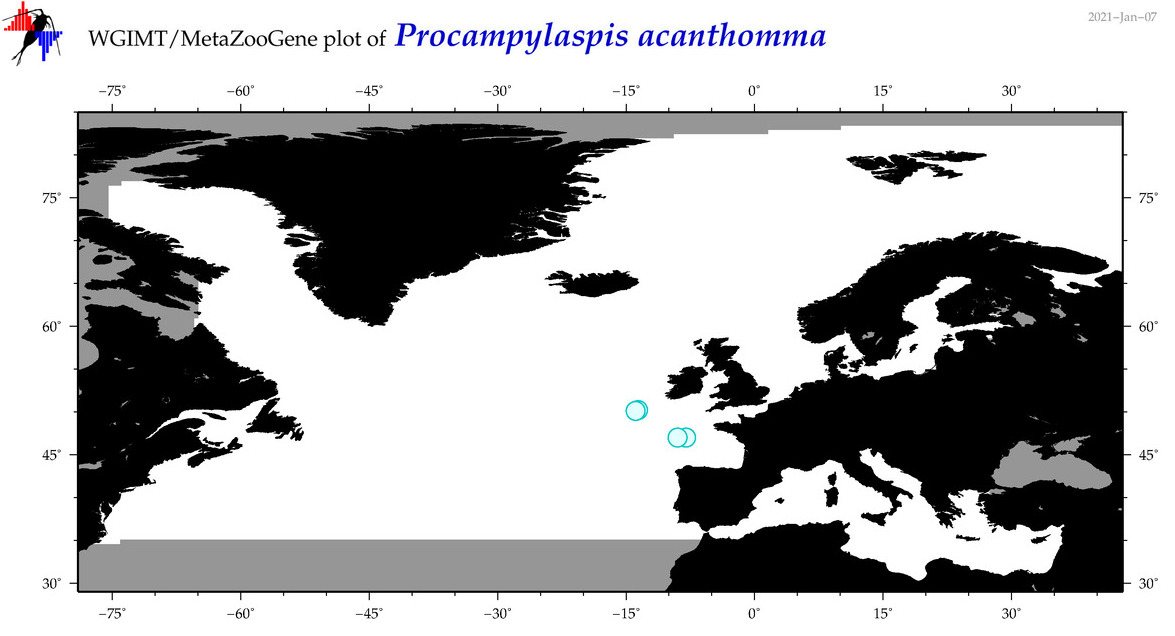
| Procampylaspis acanthomma |
Species
(118) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065352
R:1:0:0:0 |
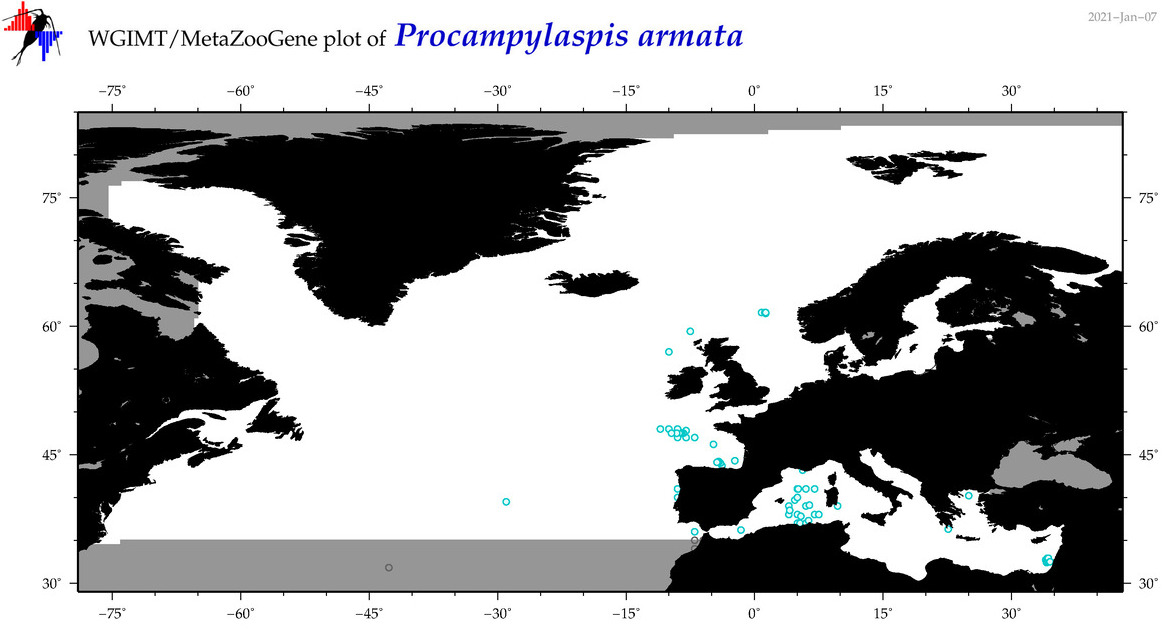
| Procampylaspis armata |
Species
(119) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065355
R:1:0:0:0 |
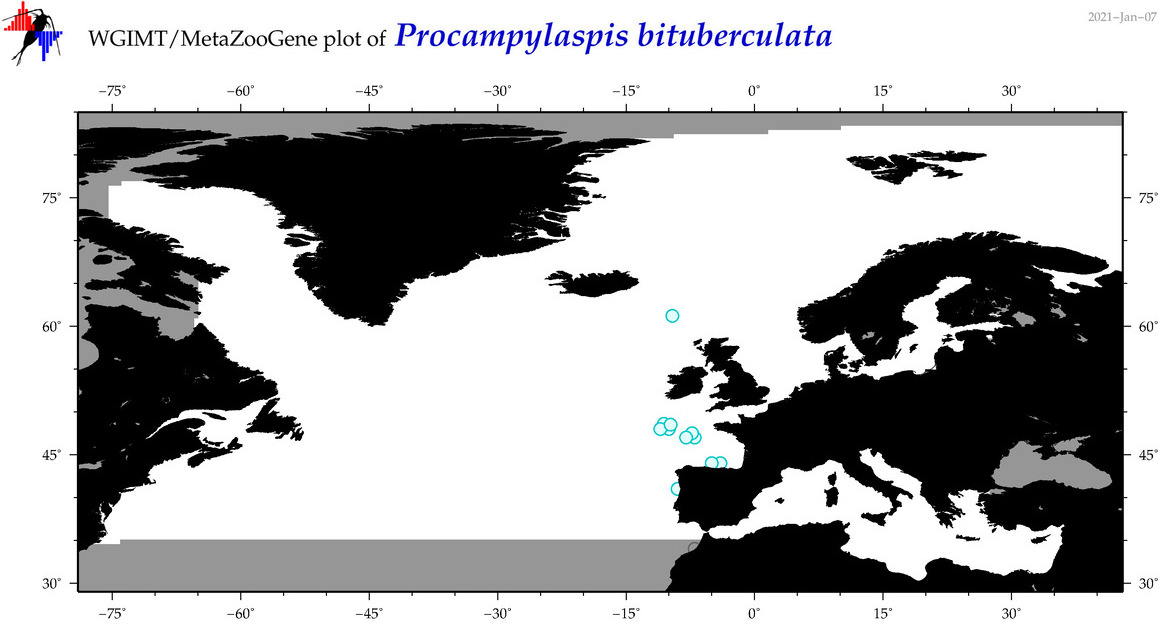
| Procampylaspis bituberculata |
Species
(120) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065359
R:1:0:0:0 |
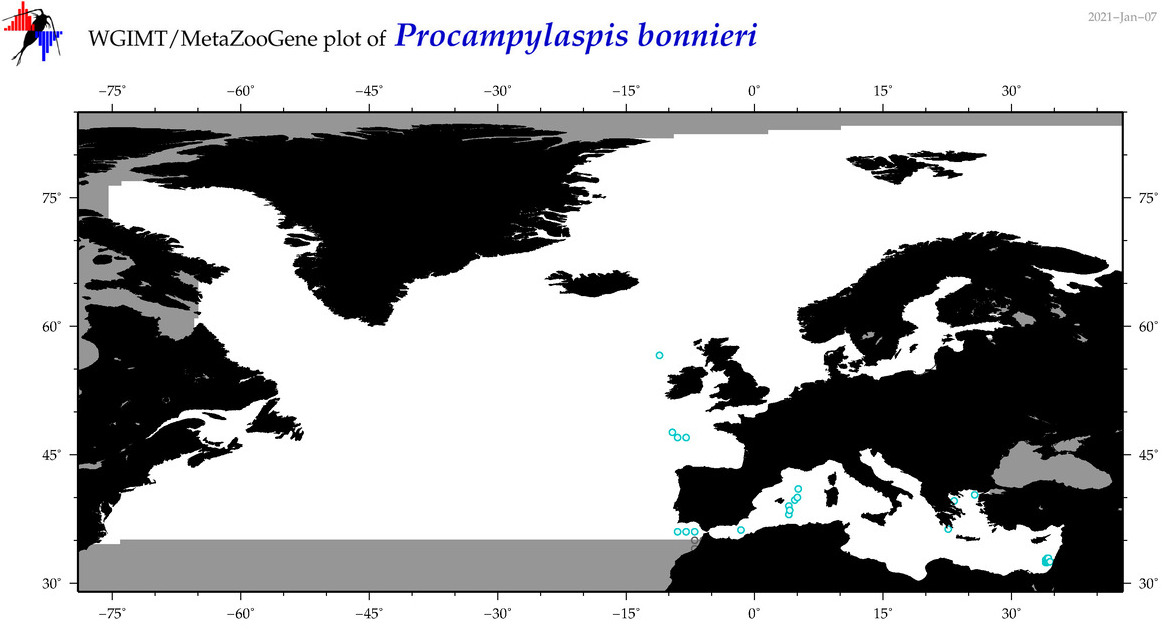
| Procampylaspis bonnieri |
Species
(121) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065360
R:1:0:0:0 |
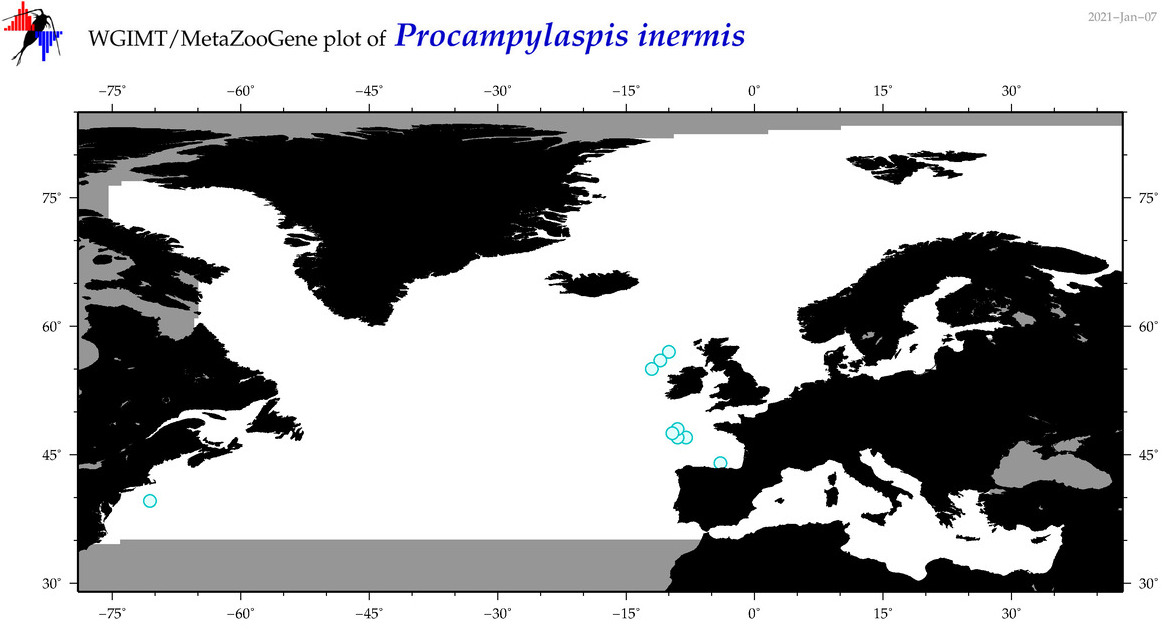
| Procampylaspis inermis |
Species
(122) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065370
R:1:0:0:0 |
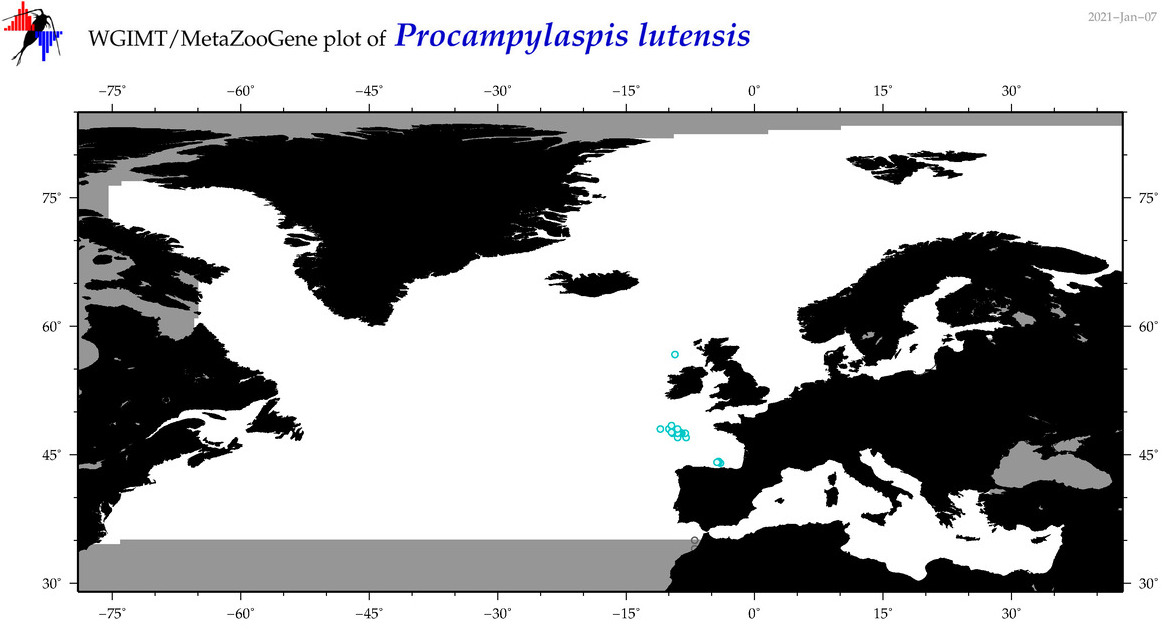
| Procampylaspis lutensis |
Species
(123) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065373
R:1:0:0:0 |
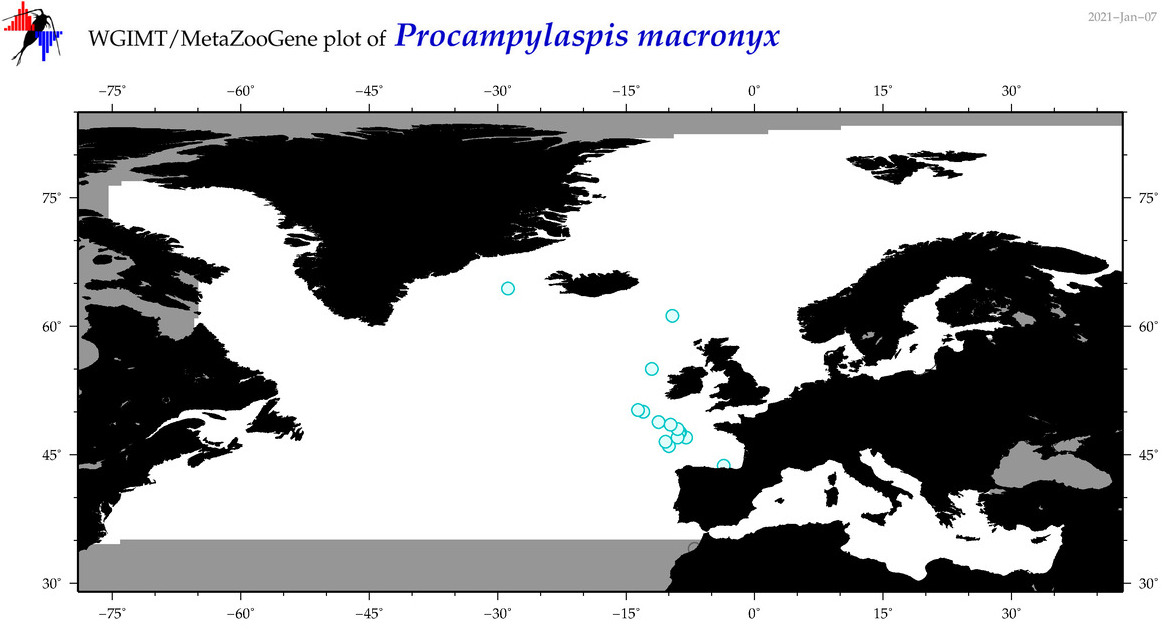
| Procampylaspis macronyx |
Species
(124) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065374
R:1:0:0:0 |
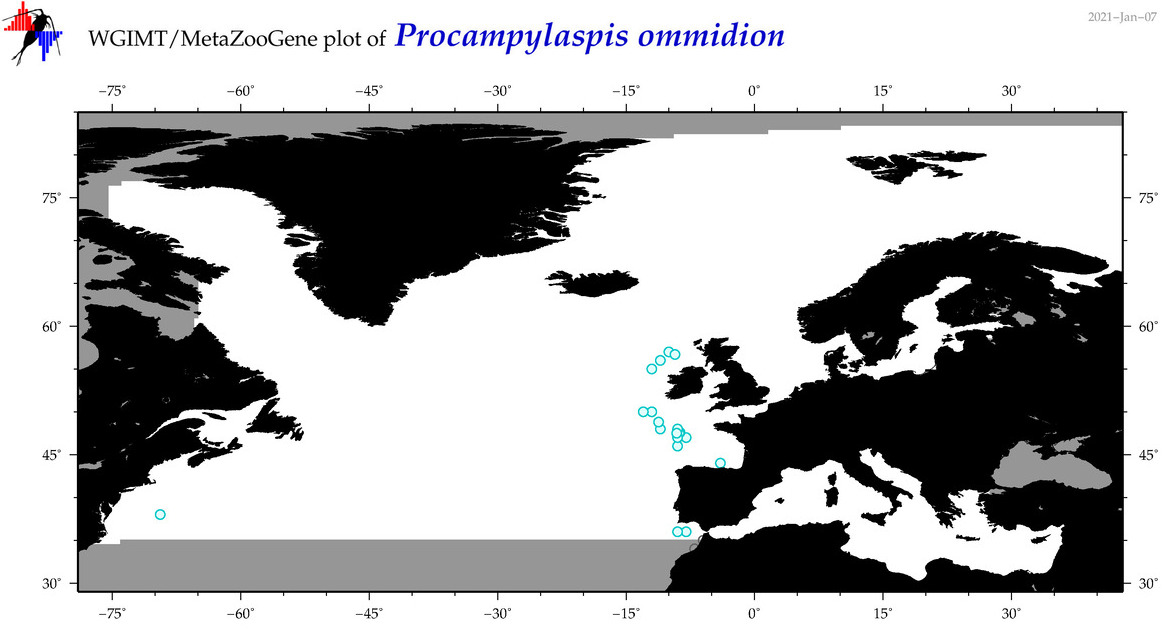
| Procampylaspis ommidion |
Species
(125) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4065376
R:1:0:0:0 |
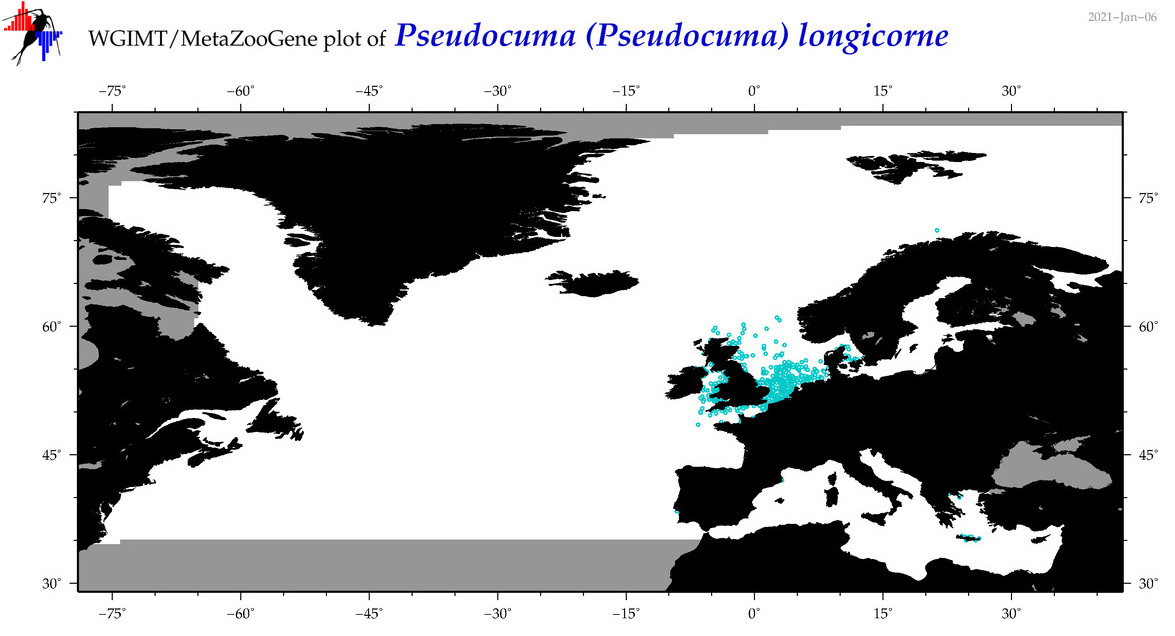
| Pseudocuma (Pseudocuma) longicorne |
Species
(126) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091502
R:1:0:0:0 |
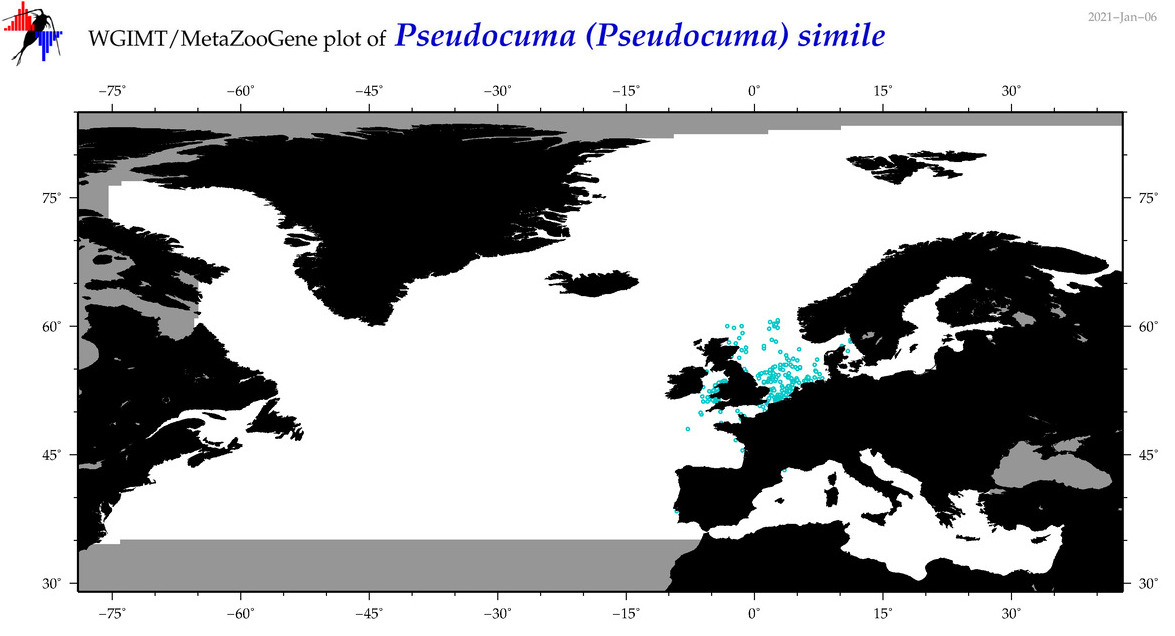
| Pseudocuma (Pseudocuma) simile |
Species
(127) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4091503
R:1:0:0:0 |
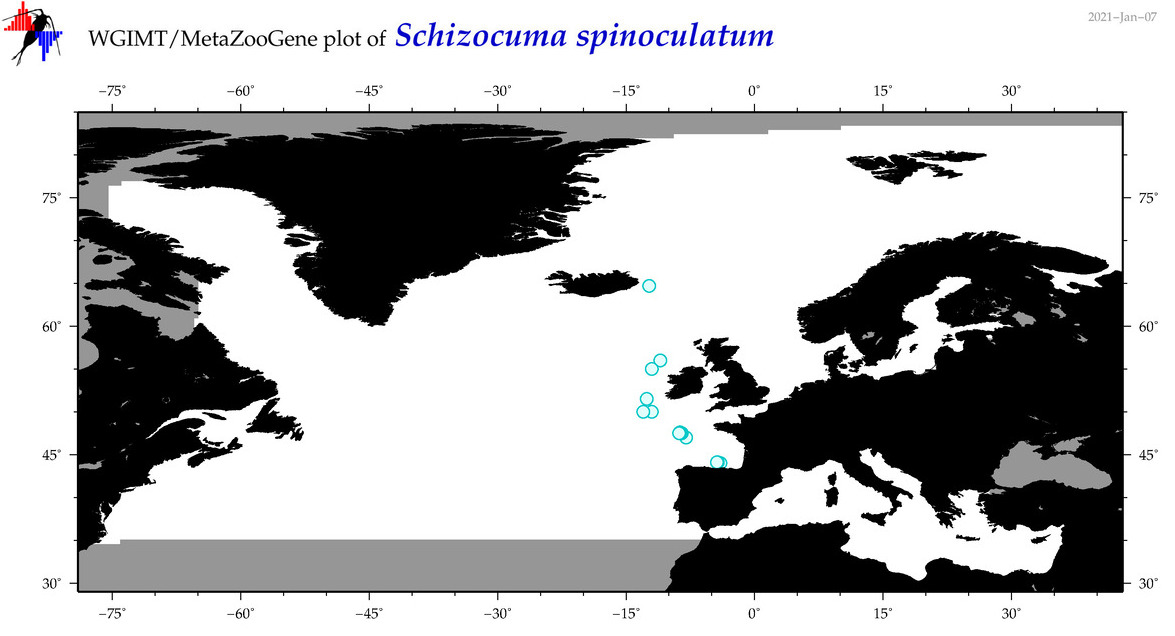
| Schizocuma spinoculatum |
Species
(128) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4067382
R:1:0:0:0 |
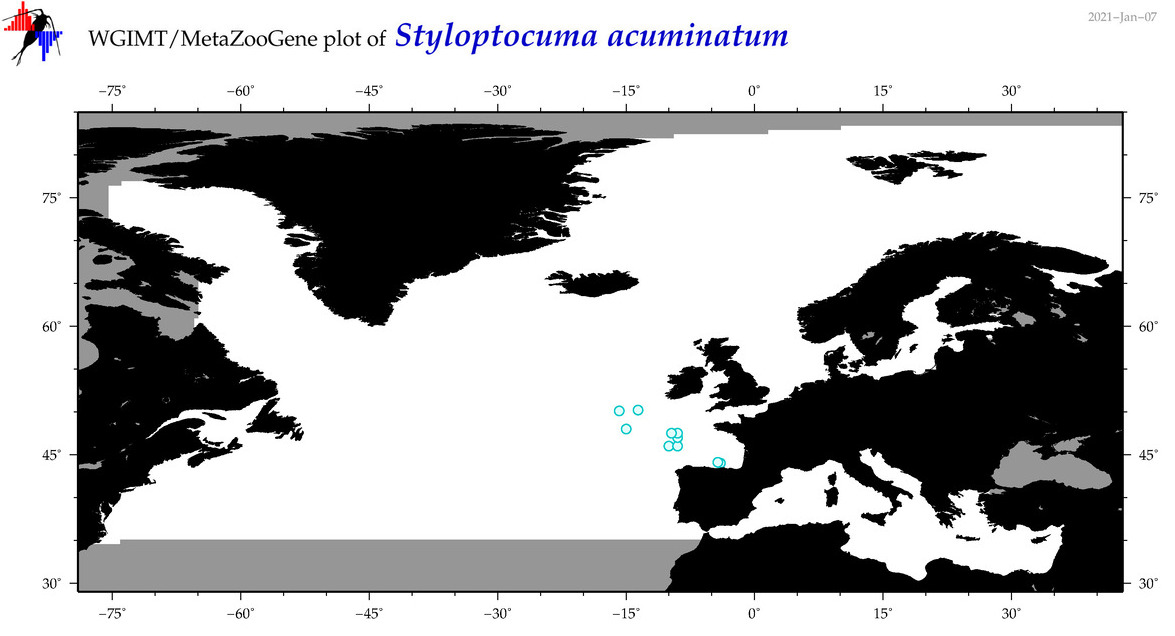
| Styloptocuma acuminatum |
Species
(129) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4069383
R:1:0:0:0 |
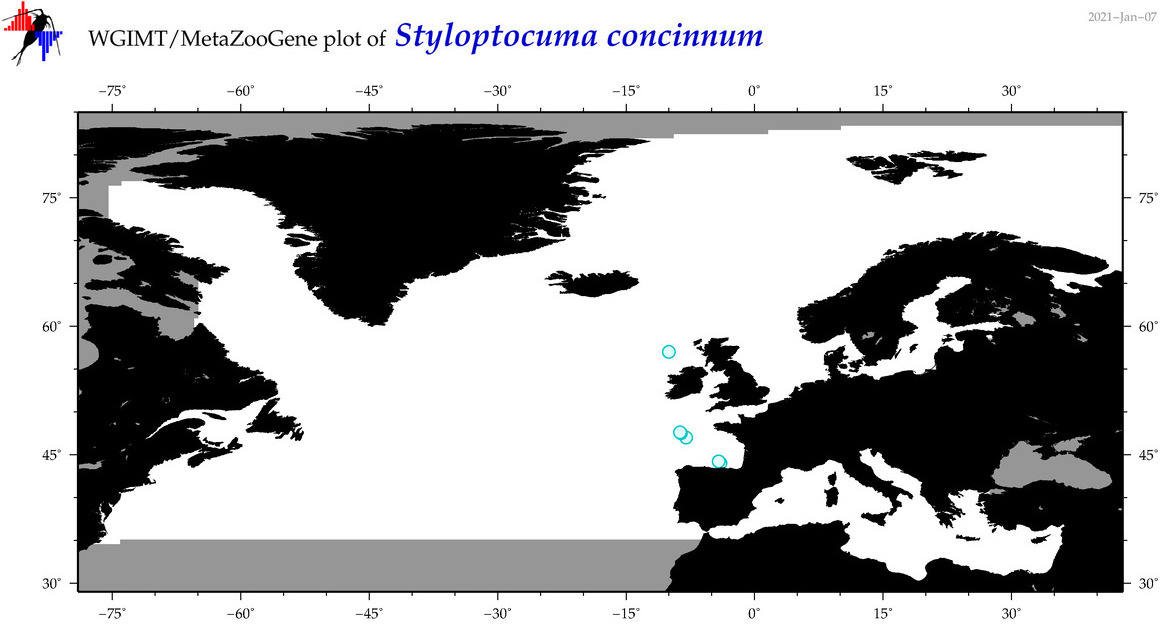
| Styloptocuma concinnum |
Species
(130) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4069387
R:1:0:0:0 |
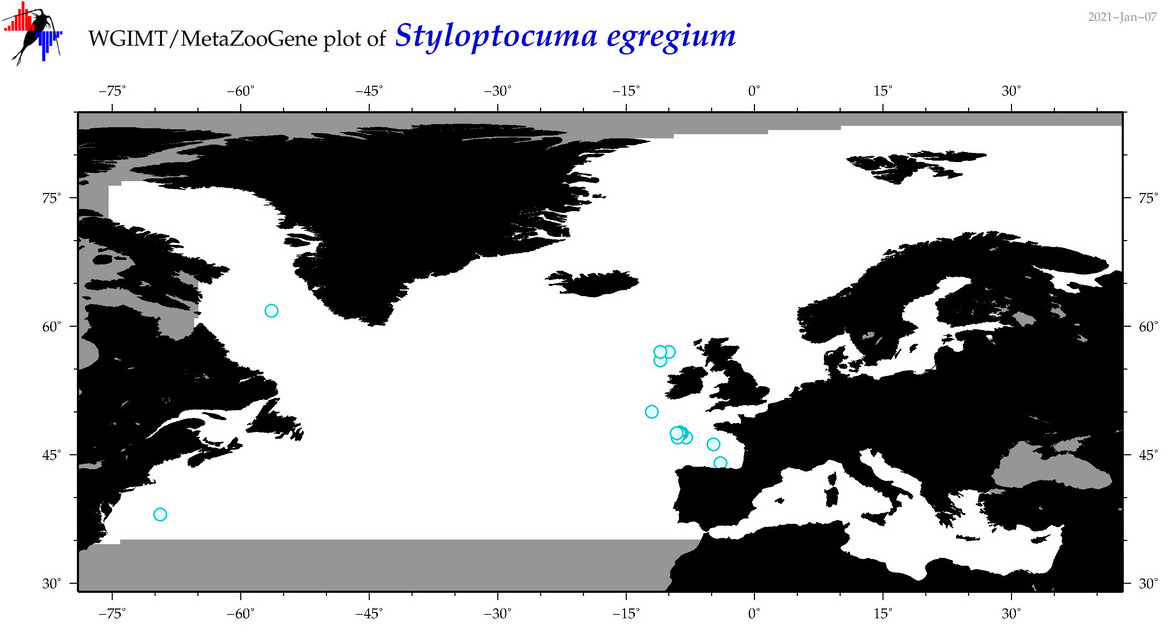
| Styloptocuma egregium |
Species
(131) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4069391
R:1:0:0:0 |
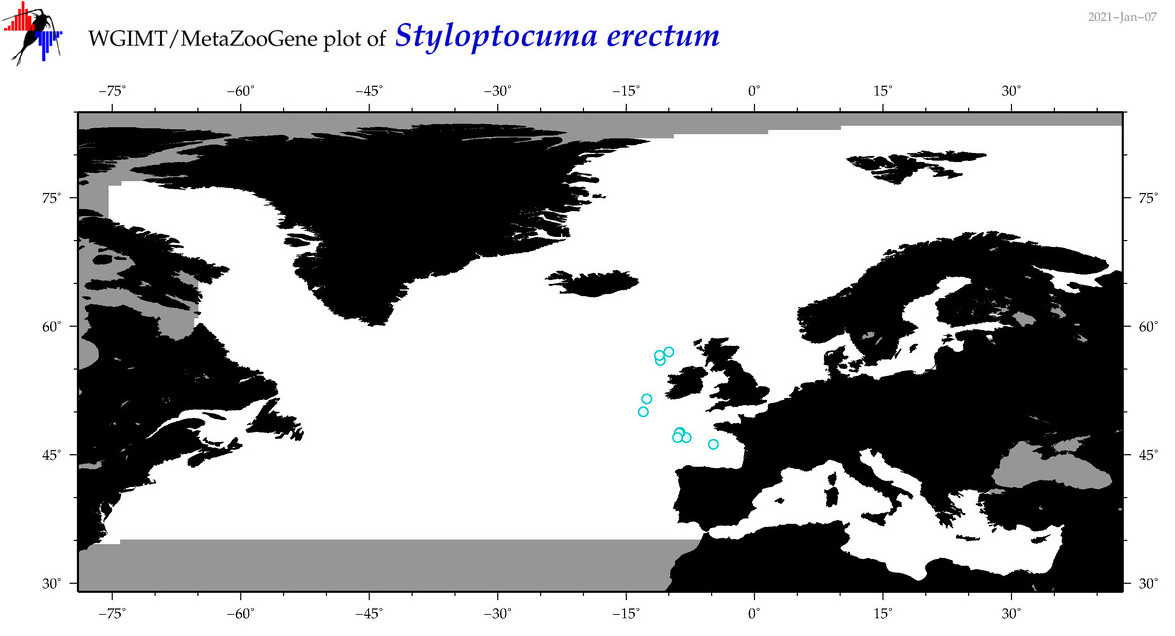
| Styloptocuma erectum |
Species
(132) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4069392
R:1:0:0:0 |
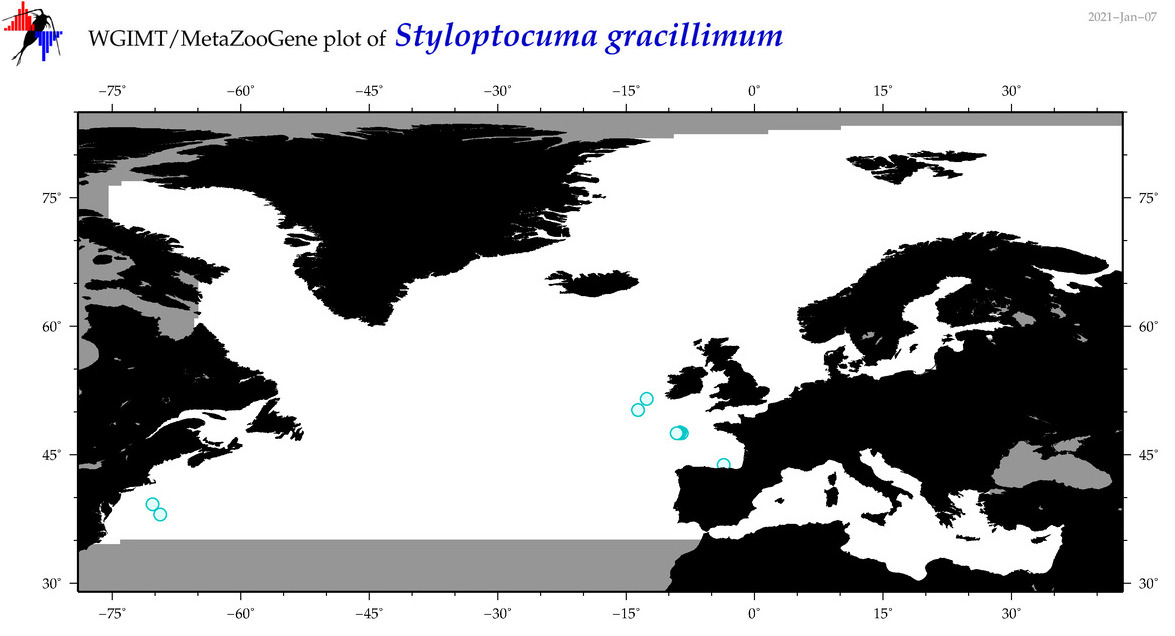
| Styloptocuma gracillimum |
Species
(133) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4069396
R:1:0:0:0 |
| Vaunthompsonia caeca |
Species
(134) |
No barcodes? |
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
no map |
accepted
T4071383
R:1:0:0:0 |
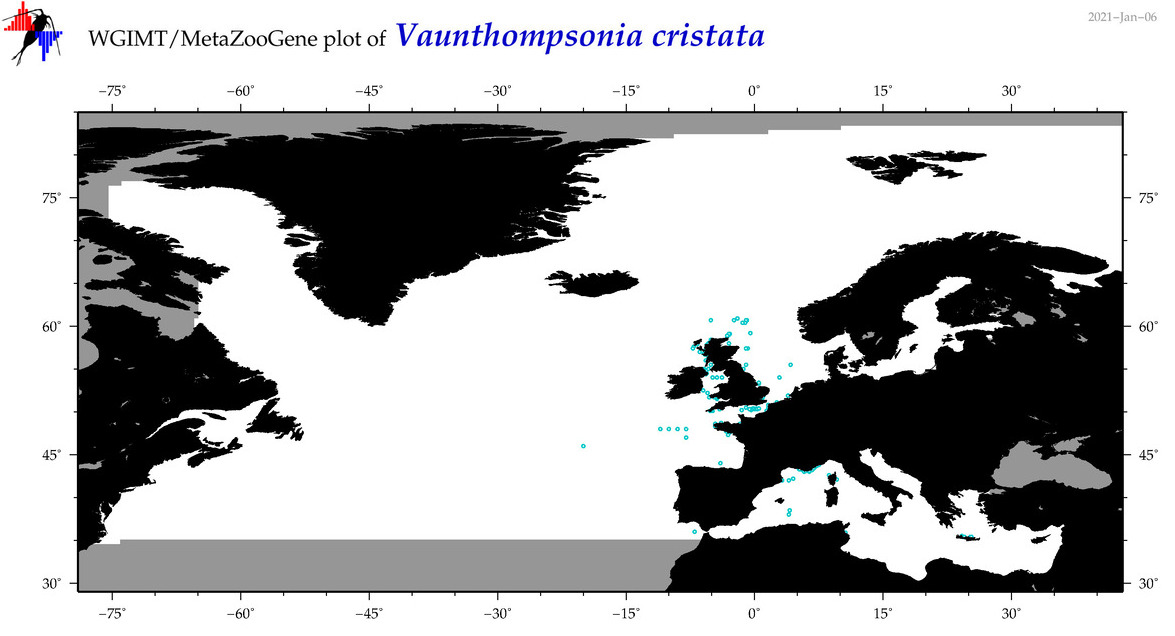
| Vaunthompsonia cristata |
Species
(135) |
COI
12S
16S
18S
28S
|
COI = 1
12S = 0
16S = 0
18S = 0
28S = 0
|
COI = 0
12S = 0
16S = 0
18S = 0
28S = 0
|
 |
accepted
T4022782
R:1:0:0:0 |